RNA-seq profiling of Pseudomonas aeruginosa under anoxic cystic fibrosis-like growth
This study uses RNA-seq to compare planktonic, short-term anoxic, and long-term anoxic biofilm states of Pseudomonas aeruginosa PA14 in synthetic cystic fibrosis sputum medium, revealing transcriptomic changes linked to denitrification, chronic adaptation, and antibiotic tolerance.
This paper moves away from RNA structure and into a very different biological setting: transcriptome adaptation of Pseudomonas aeruginosa in a cystic fibrosis-like environment. The problem is clinically important and mechanistically rich. In the lungs of cystic fibrosis patients, P. aeruginosa often encounters oxygen-limited mucus and establishes persistent biofilm-associated infections. That means the interesting question is not just how the bacterium behaves in standard aerobic culture, but how its gene-expression program changes when it transitions into sustained anoxic growth under conditions that better resemble the host environment.
That is the motivation for this study. We grew the clinical isolate PA14 in synthetic cystic fibrosis sputum medium and compared three states by RNA-seq: planktonic aerobic growth, an early response 30 minutes after the shift to anaerobiosis, and mature anoxic biofilm growth after 96 hours. This design matters because it separates acute oxygen limitation from the later, more stable transcriptional program associated with long-term adaptation. In other words, the paper is not only about “anoxia versus oxygen”, but about the trajectory from short-term response to chronic biofilm physiology.
Methodologically, this was a straightforward but strong transcriptomics experiment for its time. Directional RNA-seq libraries were prepared from biological replicates after rRNA depletion, sequenced on an Illumina platform, and mapped against the PA14 reference genome. The resulting differential-expression profiles were then interpreted at both the individual-gene and pathway level. What makes the paper useful is that it does not stop at a catalog of changed transcripts. It follows the transcriptome with mutant screening and targeted validation for selected genes, which helps connect the sequencing result back to biofilm phenotypes and antibiotic-related traits.
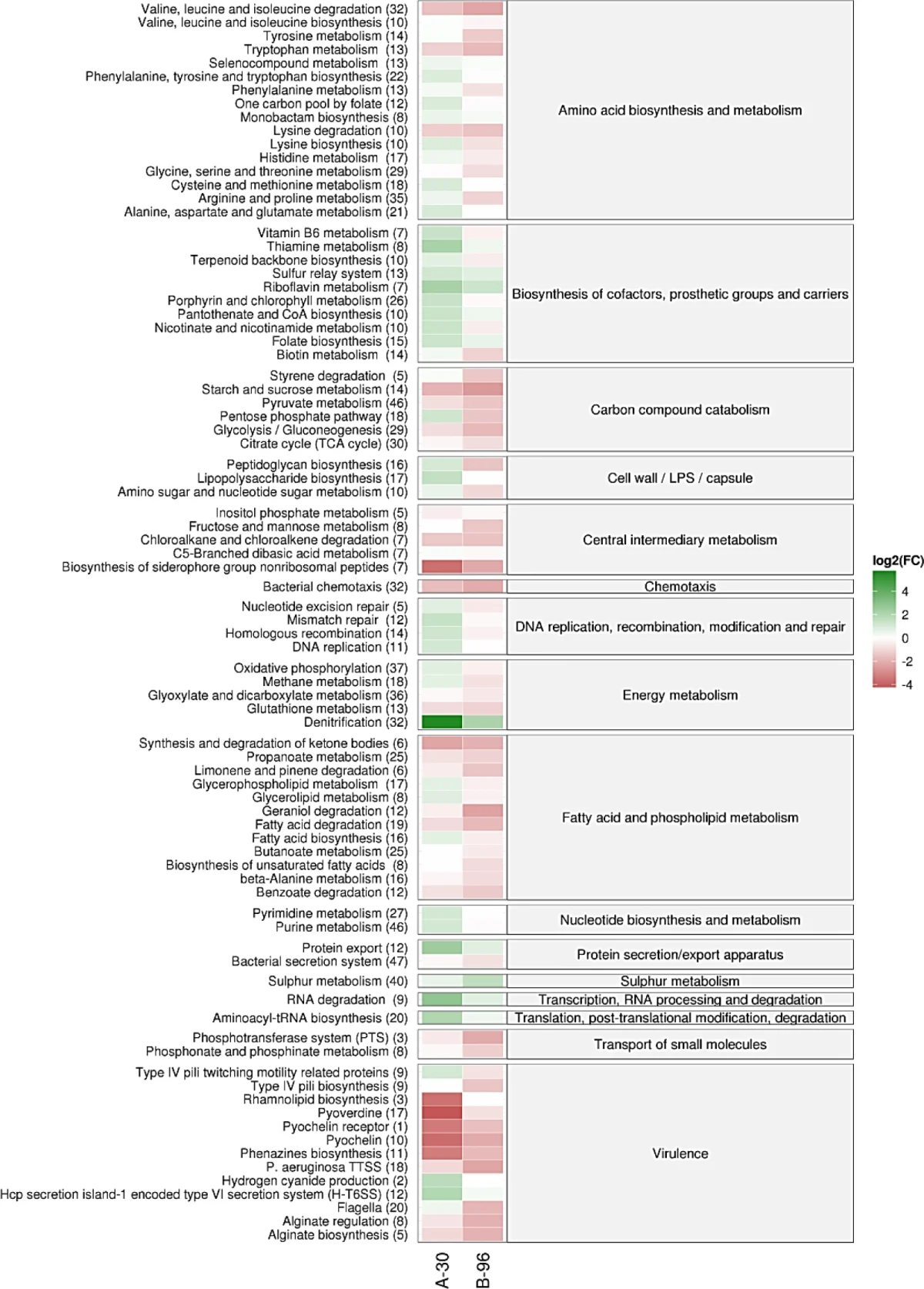
The main biological picture is clear. Short-term anoxia and long-term anoxic biofilm growth are not interchangeable states. Prolonged biofilm growth in the cystic-fibrosis-like medium is accompanied by broad pathway-level rewiring, including strong changes in denitrification and sulfur metabolism, shifts in central carbon metabolism, altered expression of virulence-associated functions, and changes in membrane or envelope-related pathways. The pathway heatmap from the paper captures this nicely: by the time the culture reaches the 96-hour biofilm state, the transcriptome has moved well beyond the immediate oxygen-starvation response.
One of the more practically relevant findings concerns antibiotic tolerance. The study reports decreased abundance of oprD transcripts and increased abundance of the mexCD-oprJ efflux operon in long-term anoxic biofilms. That combination is mechanistically interesting because it offers a plausible transcriptional explanation for altered susceptibility patterns under these chronic-growth conditions. The anoxic biofilm state does not just affect metabolism. It also reshapes how the bacterium may respond to antimicrobial pressure.
The study also identified candidate functions required for sustained anoxic biofilm growth through follow-up mutant analysis. That part is important because it turns the paper from a descriptive RNA-seq survey into a resource for bacterial physiology. Transcriptome profiling is most useful when it helps prioritize which genes or pathways deserve closer functional study, and this paper does that well in the context of chronic Pseudomonas adaptation.
From today’s perspective, the paper is an early example of a style of infection-related transcriptomics that has become standard: move closer to the host-relevant environment, distinguish transient from persistent states, and interpret gene-expression change in terms of physiology rather than as an undifferentiated list of up- and downregulated genes. For P. aeruginosa, that is especially important because oxygen limitation, nitrate respiration, biofilm formation, and drug tolerance are deeply entangled in chronic infection.
Rewiring of Gene Expression in Pseudomonas aeruginosa During Diauxic Growth Reveals an Indirect Regulation of the MexGHI-OpmD Efflux Pump by Hfq continues the same general theme from a different angle. Transcriptome-level changes in Pseudomonas are often easiest to understand when metabolism, physiology, and antibiotic response are considered together. RNA-Seq Based Transcriptional Profiling of Pseudomonas Aeruginosa Pa14 After Short- and Long-Term Anoxic Cultivation in Synthetic Cystic Fibrosis Sputum MediumCitation
Muralidhar Tata, Michael T. Wolfinger, Fabian Amman, Nicole Roschanski, Andreas Dötsch, Elisabeth Sonnleitner, Susanne Häussler, Udo Bläsi
PLoS ONE 11 (1): e0147811 (2016) | doi:10.1371/journal.pone.0147811 | PDF