The MazF regulon and post-transcriptional stress adaptation in Escherichia coli
This paper uses Poly-seq to define the MazF regulon in Escherichia coli, showing how MazF reshapes both mRNA processing and ribosome specificity to reprogram translation during harsh stress.
This paper tackles a question that sits at the intersection of RNA biology, bacterial stress physiology, and translation control: what exactly does the toxin MazF do to the cell’s gene-expression program once it is activated under stress? MazF is often introduced as part of a toxin-antitoxin system, but that label alone does not explain the biology. The more interesting issue is whether MazF simply shuts translation down indiscriminately, or whether it creates a more selective post-transcriptional program that helps cells adapt to harsh conditions.
The central claim of this study is that MazF does not merely destroy RNA in a nonspecific way. Instead, it reprograms translation. MazF cleaves a defined subset of mRNAs and also processes the 16S rRNA within mature ribosomes, removing the anti-Shine-Dalgarno region and thereby generating what had previously been described as stress-ribosomes. Those remodeled ribosomes are no longer equivalent to canonical ribosomes: they become better suited to translate MazF-processed transcripts. That immediately turns the problem into a systems-level one. If both the messages and the decoding machinery are being changed, then the relevant output is not just the transcriptome, but the translatome.
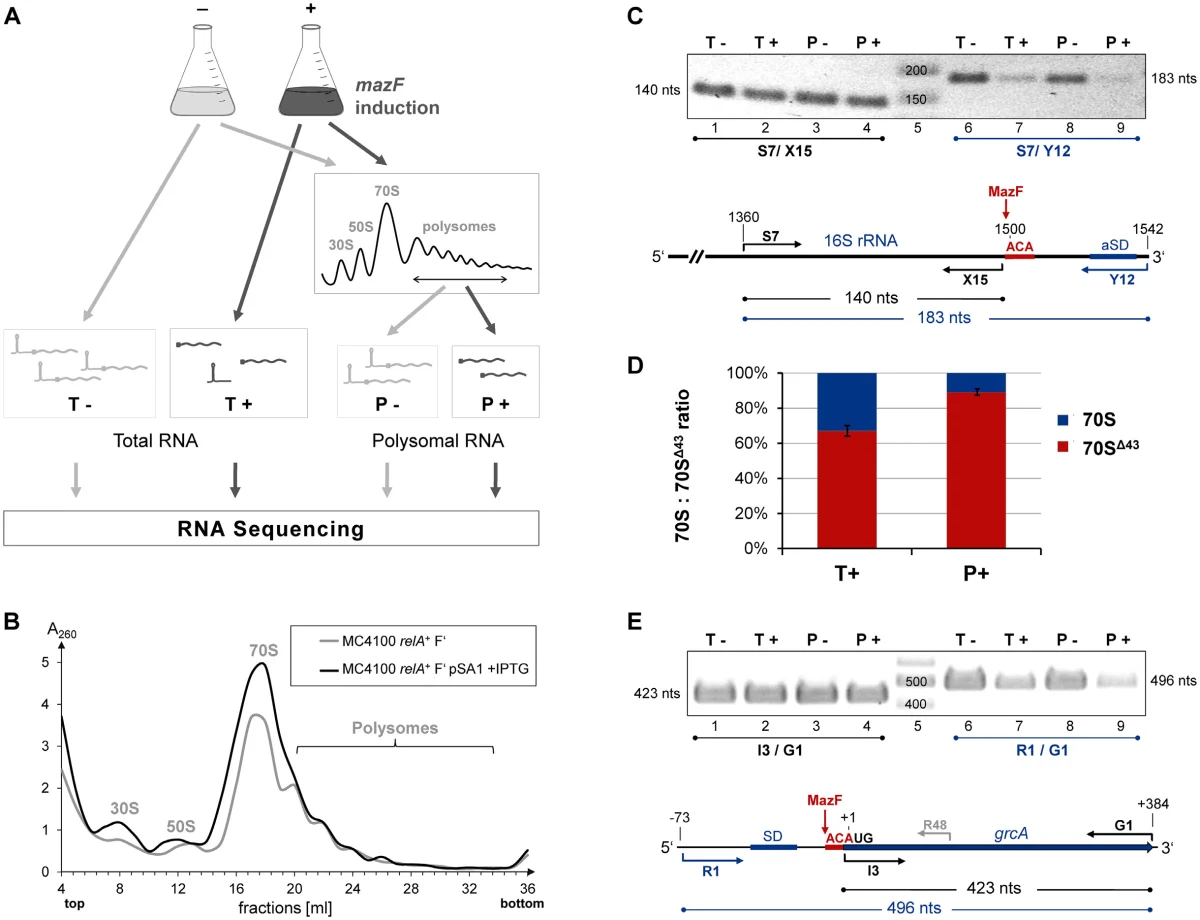
That is where the methodological contribution of the paper comes in. To define this MazF-dependent regulatory layer, we used Poly-seq, combining polysome fractionation with RNA-seq. The point of the method is that it captures intact polysome-associated transcripts rather than only short protected fragments. This makes it possible to ask, in parallel, which RNAs are present, which have been processed, and which are actually associated with translating ribosomes after MazF induction. For this question, that is exactly the right scale of observation.
The results are interesting because they resist a simplistic interpretation. One might expect a dedicated stress regulon in the narrow sense, enriched mainly for classical stress-defense factors. Instead, the MazF-regulon spans a much broader range of cellular functions. The corresponding protein products are not limited to a textbook “general stress response” module. That is one of the reasons the paper is useful: it argues that MazF-dependent translational reprogramming is not just an emergency on-off switch, but a more distributed mechanism for rapidly altering what the cell is able to synthesize under severe stress.
This broader view also changes how one thinks about selective translation in bacteria. In many discussions of bacterial regulation, transcription still gets the starring role, with translational control treated as a local fine-tuning layer. MazF is a strong counterexample. Here, post-transcriptional regulation acts globally enough to reshape the functional output of the cell, and it does so on a timescale that makes sense for acute stress adaptation. The paper therefore highlights selective translation as a major regulatory mechanism in its own right rather than a secondary detail downstream of transcription.
Another important aspect is the connection to persistence. Toxin-antitoxin systems have long been discussed in relation to persister-cell formation, but the mechanistic links often remain vague. This work does not reduce that problem to a single answer, yet it makes a concrete contribution: if MazF introduces heterogeneity in which transcripts remain translation-competent and which ribosomes are available to decode them, then it provides a plausible route by which a stressed bacterial population can diversify its phenotypic state. That makes MazF a candidate effector of harsh-stress adaptation rather than a passive marker of it.
From my perspective, one of the strengths of this paper is that it combines a fairly elegant conceptual model with a method that can actually test it. The figure from the paper captures the logic well: MazF induction, separation of total and polysome-associated RNA, sequencing-based identification of processed transcripts, and direct evidence for MazF-dependent remodeling of the ribosome itself. That is what turns the “MazF regulon” from a loose idea into something experimentally definable.
So although the title uses the word “toolbox”, the main message is not simply that MazF controls a list of genes. It is that bacteria can implement a fast post-transcriptional stress response by simultaneously editing the message pool and the translation machinery. For anyone interested in bacterial RNA biology, translational control, or stress-induced phenotypic diversification, that remains a useful framework. The MazF-Regulon: A Toolbox for the Post-Transcriptional Stress Response in Escherichia ColiCitation
Martina Sauert, Michael T. Wolfinger, Oliver Vesper, Christian Müller, Konstantin Byrgazov, Isabella Moll
Nucleic Acids Res. 44 (14): 6660–6675 (2016) | doi:10.1093/nar/gkw115 | PDF