Chikungunya virus phylogeny and lineage-specific RNA structures
An updated CHIKV phylogeny based on 598 genomes, linking lineage structure to conserved and lineage-specific 3' UTR RNA architectures.
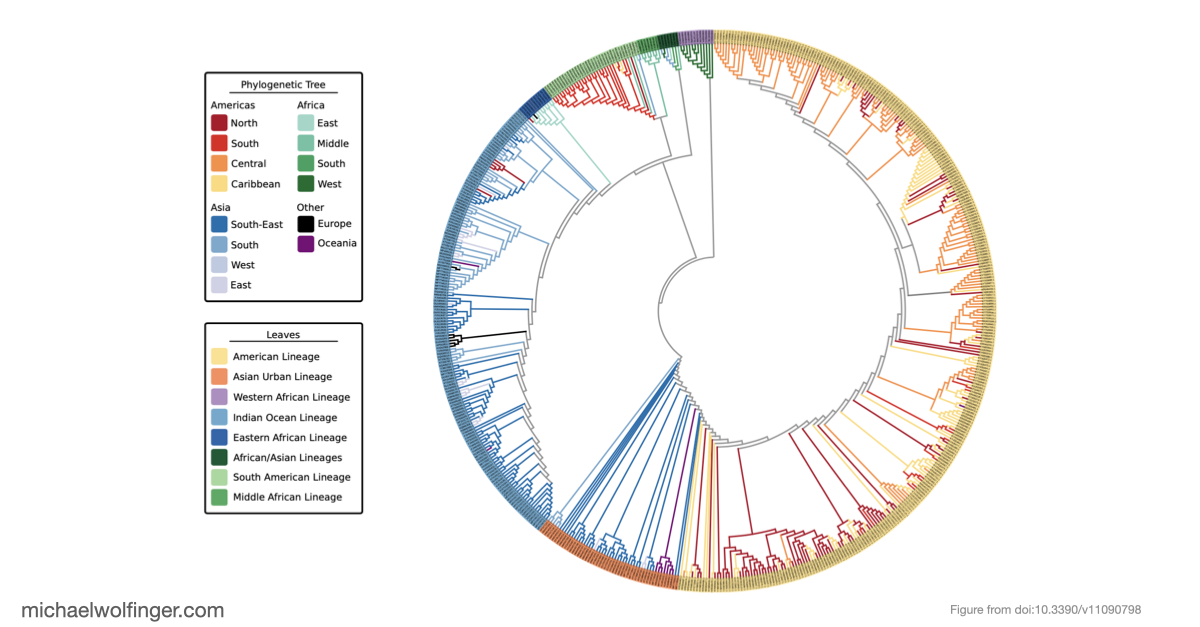
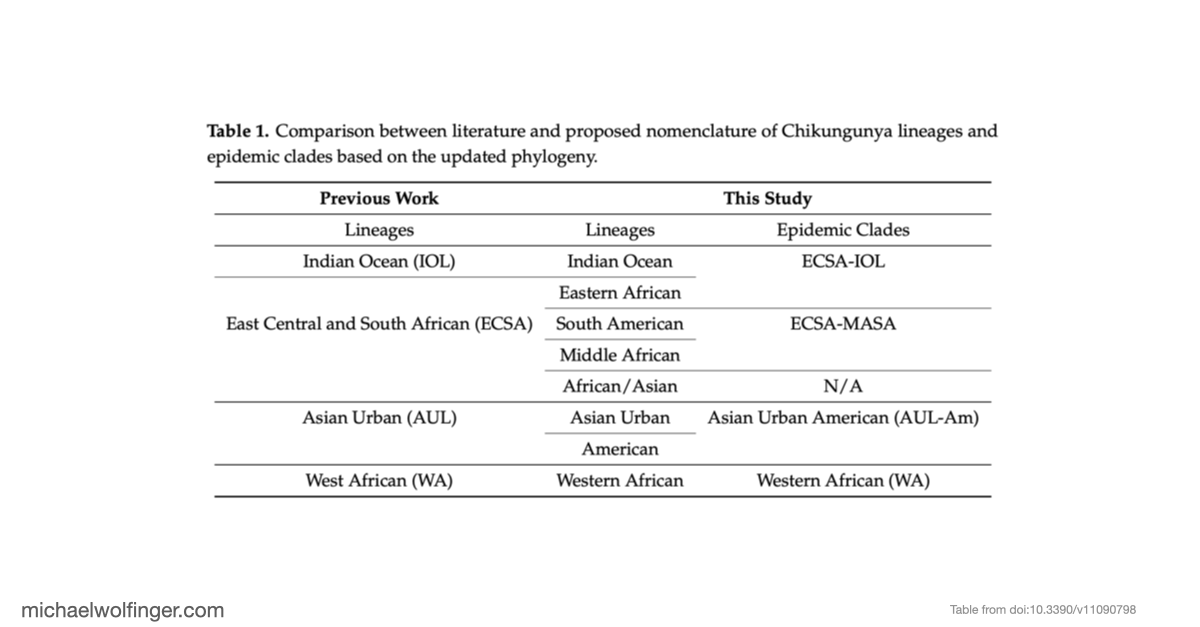
Chikungunya virus (CHIKV) is usually discussed in terms of a small set of named geographic lineages, but that picture has become increasingly inadequate as more genome sequences have accumulated. In this study we revisited CHIKV evolution using 598 complete genomes and asked two connected questions: how well the traditional lineage labels still reflect the phylogeny, and whether the distinct clades also differ in the organization of their untranslated regions.
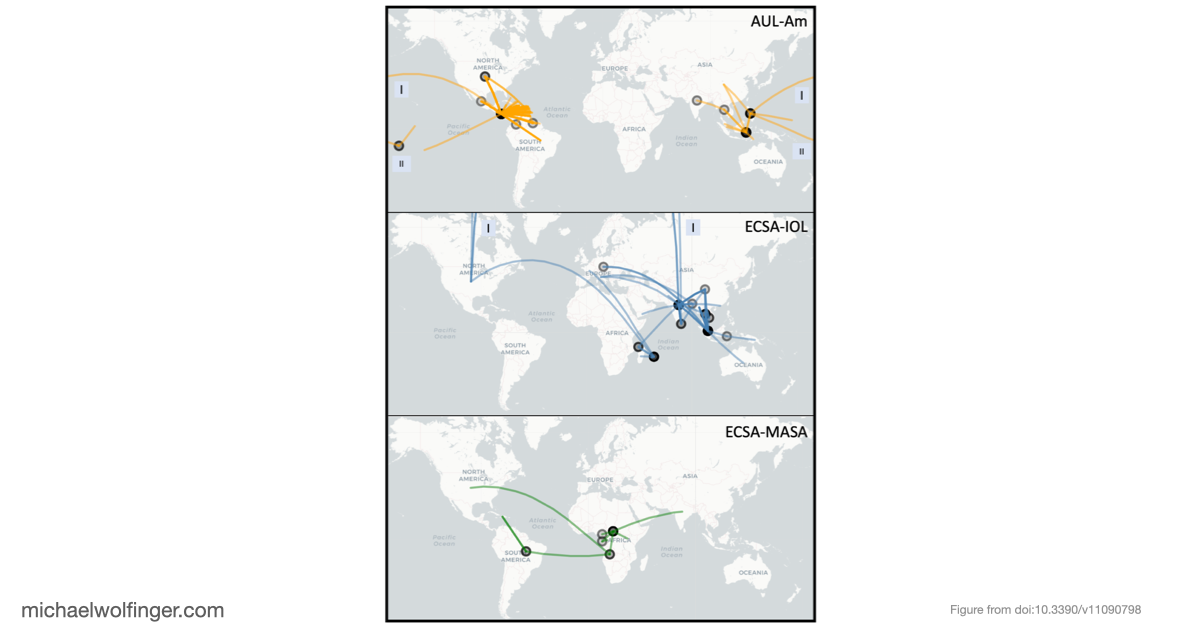
The phylogenetic analysis was based on the two viral open reading frames and used a maximum-likelihood framework with explicit support assessment. One of the central results is that the historically defined ECSA group is not best understood as a single lineage. Instead, the data support several African sublineages with different evolutionary histories, including Eastern, Middle, and Southern African components. That distinction matters because the major epidemic clades outside Africa can be traced back to different parts of this broader African diversity. In the revised view used in the paper, the Indian Ocean lineage derives from Eastern African ancestors, while the South American outbreak strain is linked to Middle African taxa.
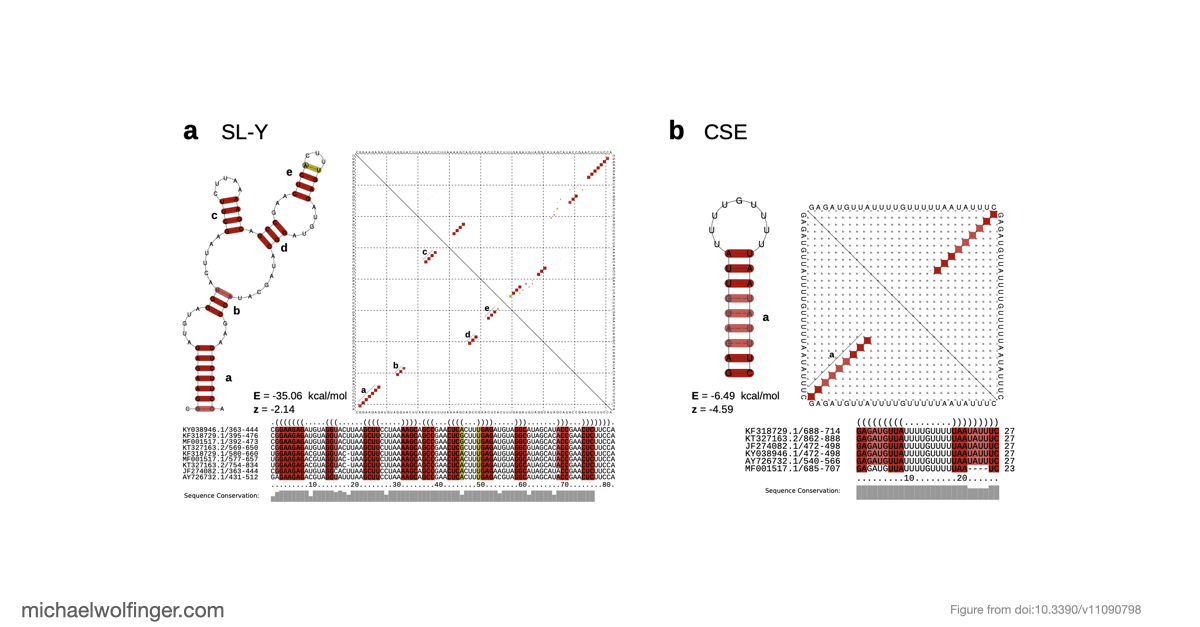
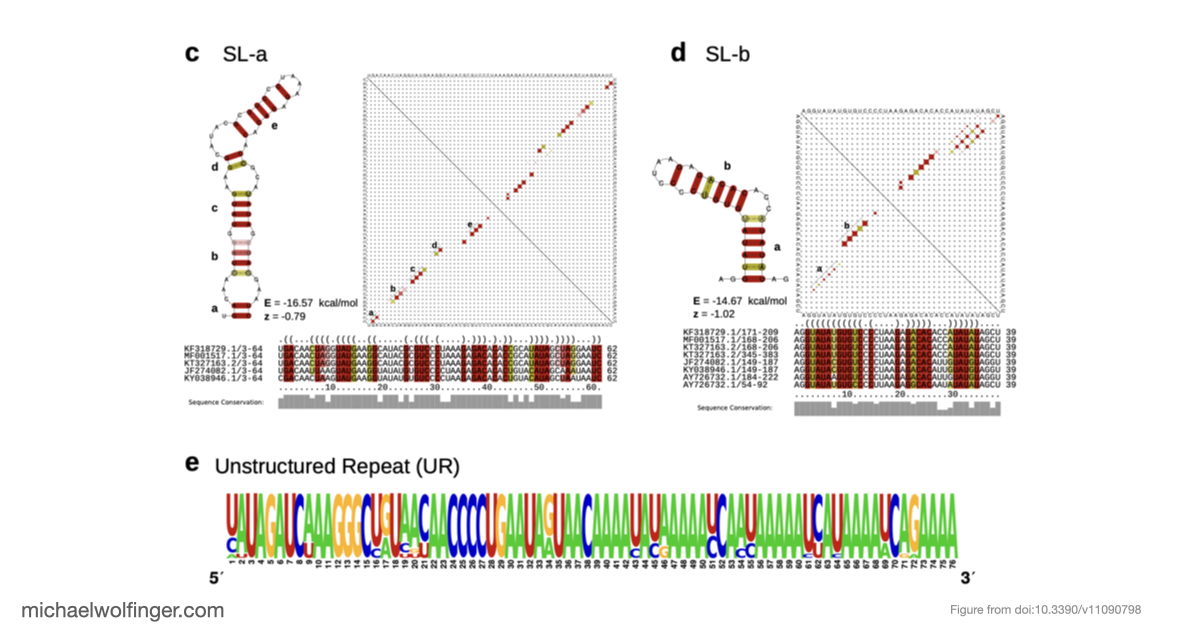
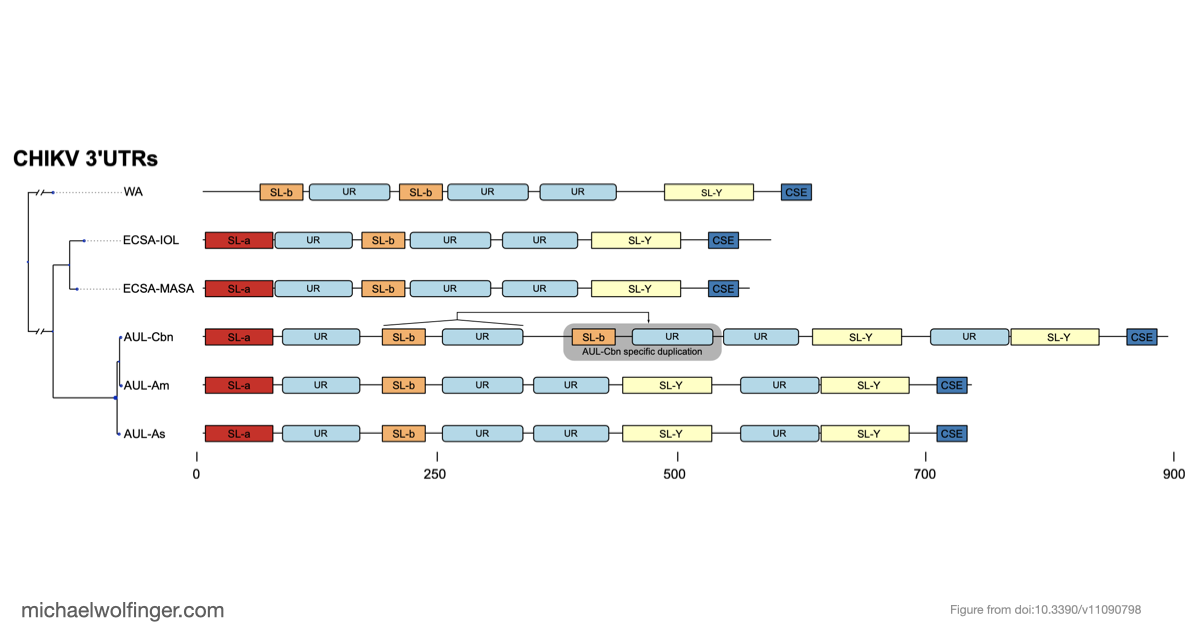
The second part of the paper focuses on the CHIKV 3' UTR, a region known to carry repeated sequence elements but whose structural organization had remained unclear. Using comparative RNA analysis, covariance-model based homology searches, and consensus secondary structure prediction, we found that the repeat-rich 3' UTR is not just a collection of duplicated sequence blocks. Instead, it follows an alternating architecture of conserved structured and unstructured elements. The structured part consists of stem-loop elements termed SL-a and SL-b, a Y-shaped element SL-Y, and a conserved sequence element near the terminus, while the intervening repeat regions remain predominantly unpaired.
This structured-versus-unstructured organization is one of the more interesting aspects of the study. The unstructured repeat regions are unlikely to be mere spacers. Because they remain accessible, they may act as structural insulators between neighboring RNA elements or provide binding surfaces for host proteins. The paper points in particular to repeated UAG motifs in these accessible regions, which makes interactions with proteins such as Musashi plausible, although that remains a functional hypothesis rather than a demonstrated mechanism.
Another useful outcome is that the 3' UTR architectures differ systematically between lineages. Some elements are shared across essentially all CHIKV groups, while others vary in copy number or arrangement, and recent American isolates of the Asian Urban lineage show evidence of partial duplication events. This suggests that CHIKV evolution in the 3' UTR is not just sequence drift. Rather, the virus appears to rearrange modular RNA building blocks while preserving an overall architectural pattern. That is a more informative model than treating the repeat region as a simple low-complexity tail.
From a computational RNA perspective, this paper is also a good example of what comparative genomics can do even when strong covariation support is limited. CHIKV 3' UTR sequences are often too similar, and the available sampling is too outbreak-biased, to expect the kind of covariation signal seen in deeply diverged structured RNAs. Even so, combining phylogeny, repeated architecture, local thermodynamic stability, and homology modeling is enough to identify plausible conserved RNA elements and to formulate experimentally testable hypotheses about their role in replication, host adaptation, and lineage-specific biology. Updated Phylogeny of Chikungunya Virus Suggests Lineage-Specific RNA Architecture Dynamic Molecular Epidemiology Reveals Lineage-Associated Single-Nucleotide Variants That Alter RNA Structure in Chikungunya Virus Figures and Data
Citation
Adriano de Bernardi Schneider, Roman Ochsenreiter, Reilly Hostager, Ivo L. Hofacker, Daniel Janies, Michael T. Wolfinger
Viruses 11:798 (2019) | doi:10.3390/v11090798 | PDF | FiguresSee Also
Thomas Spicher, Markus Delitz, Adriano de Bernardi Schneider, Michael T. Wolfinger
Genes 12 (2):239 (2021) | doi:10.3390/genes12020239 | PDF | Figures