Insect-specific flaviviruses from Zambia and their exoribonuclease-resistant RNAs
This study isolates two insect-specific flaviviruses from mosquitoes in Zambia and shows that their 3'UTRs contain functional xrRNA-like elements that stall Xrn1.
This paper sits at a point where field virology, genome sequencing, and comparative RNA analysis come together very directly. The biological starting point is straightforward: mosquito populations in Zambia were screened for flaviviruses, and the resulting isolates were characterized at the genomic level. What makes the study especially interesting from my perspective is that it does not stop at virus discovery. It follows through into the untranslated regions and asks whether the newly isolated genomes also carry the conserved structured RNAs that are increasingly recognized as functional elements in flavivirus biology.
The study reports two closely related insect-specific flaviviruses from Culex mosquitoes, a Barkedji virus isolate from Zambia and a more distinct Barkedji-like virus. Both replicate in mosquito cells but not in mammalian or avian cell lines, which places them firmly in the insect-specific part of flavivirus diversity. That host restriction is already biologically interesting, but it also makes these viruses useful comparative models. They let us ask which RNA elements belong to the deeper flaviviral toolkit and which ones track more closely with host range or vector association.
Methodologically, the paper combines several layers of analysis. Viral genomes were recovered from field isolates by next-generation sequencing and end-completion methods, then compared with known flavivirus relatives. The untranslated regions were screened with comparative genomics and RNA secondary structure prediction in order to identify canonical flaviviral building blocks in both the 5' and 3' ends. This is the part where the paper moves beyond cataloguing a new isolate: it uses structural homology, not just sequence similarity, to identify conserved functional candidates.
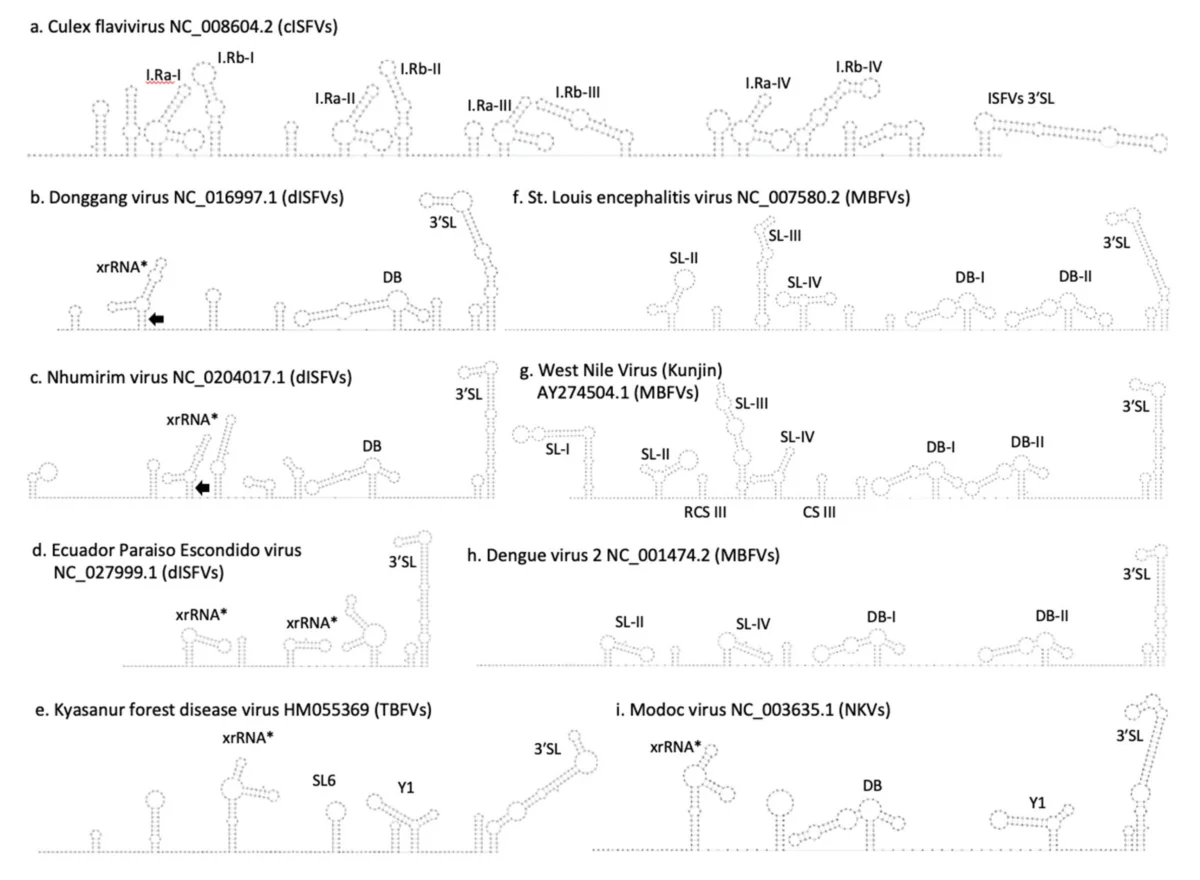
The main result is that the 3'UTRs of both Zambian isolates contain a recognizable set of structured RNA elements, including xrRNA-like folds, an SL-III element, dumbbell-like structures, and the terminal 3' stem-loop. In other words, even these insect-specific viruses retain the broader architectural logic seen across flaviviral non-coding regions. That matters because flaviviral UTRs evolve quickly at the sequence level, and purely sequence-based annotation can miss homologous elements whose function is preserved through structure rather than exact nucleotide identity.
The paper then takes the crucial extra step and tests the predicted xrRNA candidates experimentally. In vitro Xrn1 resistance assays show that the proposed elements are not just plausible folds on paper. They are able to stall the host exoribonuclease. That functional validation is important. It turns a comparative prediction into evidence that these insect-specific viruses likely generate protected decay intermediates in the same general way as other flaviviruses with established xrRNA biology.
This study helped sharpen a recurring theme in flavivirus RNA biology: structured RNAs in viral UTRs are not decorative sequence features. They are conserved control elements, and they remain informative even in relatively under-sampled corners of flavivirus diversity. The Zambia isolates expanded that picture by showing that xrRNA-associated architecture is not restricted to the most intensively studied human pathogens. It is also present in insect-specific lineages, where it can be studied in an evolutionary context that is less confounded by vertebrate pathogenicity. For a broader comparative view, this paper pairs naturally with Functional RNA Structures in the 3’UTR of Tick-Borne, Insect-Specific and No Known Vector Flaviviruses and Functional RNA Structures in the 3’UTR of Mosquito-Borne Flaviviruses.
This also makes the paper a useful bridge between virus discovery and comparative RNA biology. It starts with surveillance and isolation, but it ends with a mechanistic claim about structured non-coding RNA. That combination is one reason why insect-specific flaviviruses remain so informative: they provide natural experiments in how conserved RNA elements are maintained, duplicated, or remodeled across different ecological niches. Discoveries of Exoribonuclease-Resistant Structures of Insect-Specific Flaviviruses Isolated in Zambia Functional RNA Structures in the 3’UTR of Tick-Borne, Insect-Specific and No Known Vector Flaviviruses Functional RNA Structures in the 3’UTR of Mosquito-Borne FlavivirusesCitation
Christida E. Wastika, Hayato Harima, Michihito Sasakai, Bernard M. Hang'ombe, Yuki Eshita, Qiu Yongjin, William W. Hall, Michael T. Wolfinger, Hirofumi Sawa, Yasuko Orba
Viruses 12:1017 (2020) | doi:10.3390/v12091017 | PDFSee Also
Roman Ochsenreiter, Ivo L. Hofacker, Michael T. Wolfinger
Viruses 11:298 (2019) | doi:10.3390/v11030298 | PDF
Michael T. Wolfinger, Roman Ochsenreiter, Ivo L. Hofacker
In Virus Bioinformatics, pp65-100. Chapman and Hall/CRC Press (2021) | doi:10.1201/9781003097679-5 | PDF