Hfq regulates carbapenem susceptibility in Pseudomonas aeruginosa
This paper shows that the RNA chaperone Hfq controls carbapenem susceptibility in Pseudomonas aeruginosa through two different post-transcriptional routes acting on the porins OprD and OpdP.
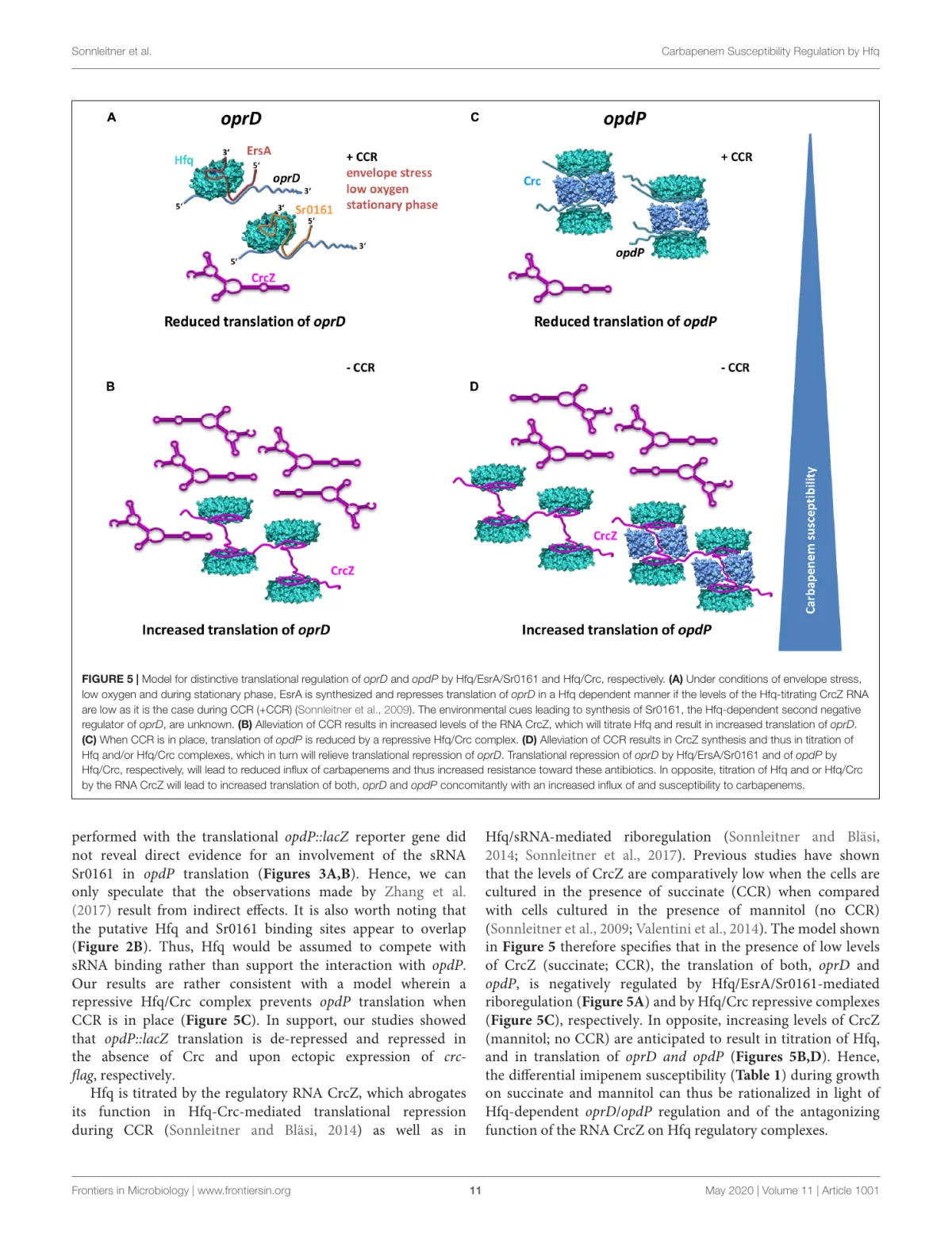
This paper addresses a clinically relevant question through a very RNA-centered mechanism. In Pseudomonas aeruginosa, carbapenem susceptibility depends in part on whether the antibiotics can enter the cell through outer-membrane porins. Two of the important entry points are OprD and OpdP. What this study shows is that these two porins are not regulated in the same way, even though both are under the influence of the RNA chaperone Hfq. That distinction matters because it links antibiotic susceptibility not just to gene expression in general, but to specific post-transcriptional circuits.
The central result is that Hfq governs oprD and opdP through different regulatory mechanisms. Translation of oprD is repressed through Hfq-dependent riboregulation involving the small RNAs ErsA and Sr0161. By contrast, opdP is not best explained by the same sRNA route. Instead, the data support direct translational repression by an Hfq/Crc complex, the same regulatory logic that is already known from carbon catabolite repression in Pseudomonas. In other words, the two carbapenem entry ports sit under related but clearly distinct post-transcriptional control systems.
Methodologically, the paper disentangles these routes with a clean genetic design. Translational lacZ reporter fusions for oprD and opdP were combined with mutant backgrounds lacking hfq, crc, ersA, or sr0161, together with complementation and ectopic expression experiments. The study also used Hfq co-immunoprecipitation and microscale thermophoresis to show that Hfq can bind relevant regions in both transcripts. That is important because it separates the question of Hfq binding from the question of which regulatory partners actually mediate repression in each case.
The mechanistic picture that emerges is appealingly specific. oprD behaves like a classic Hfq-assisted sRNA target: the porin is repressed through the action of ErsA and Sr0161, especially under conditions such as envelope stress, low oxygen, or stationary phase. opdP, on the other hand, fits better into the carbon-catabolite-repression framework, where Hfq and Crc assemble a repressive complex on target mRNAs. The paper therefore does more than identify Hfq as a regulator. It shows that one global RNA-binding protein can shape antibiotic susceptibility by deploying different regulatory strategies on different mRNA targets.
An especially useful part of the paper is the integration of CrcZ into this story. CrcZ is an Hfq-sequestering RNA whose abundance depends on carbon source and growth state. When CrcZ levels rise, Hfq and Hfq/Crc repression are relieved, which increases translation of both oprD and opdP. That gives the study a broader physiological meaning: susceptibility to carbapenems is not fixed, but can shift with metabolic state because the regulatory RNAs and RNA-protein complexes controlling porin synthesis also respond to nutrient conditions.
This makes the paper relevant beyond one antibiotic class. It is a good example of how bacterial metabolism, RNA regulation, and antimicrobial susceptibility intersect. The most interesting point is not simply that Hfq affects drug response. It is that the effect can be rationalized mechanistically, through differential control of uptake channels and through the antagonistic action of a regulatory RNA that senses the carbon regime of the cell.
This study helps define a recurring theme in Pseudomonas RNA biology: carbon catabolite repression and Hfq-dependent regulation are not abstract layers sitting above physiology. They have direct consequences for traits that matter clinically, including biofilm behavior, nutrient adaptation, and susceptibility to antibiotics. This 2020 paper is one of the clearest demonstrations of that connection at the level of individual uptake systems. It also connects naturally to Interplay Between the Catabolite Repression Control Protein Crc, Hfq and RNA in Hfq-Dependent Translational Regulation in Pseudomonas aeruginosa and to Rewiring of Gene Expression in Pseudomonas aeruginosa During Diauxic Growth Reveals an Indirect Regulation of the MexGHI-OpmD Efflux Pump by Hfq. Distinctive Regulation of Carbapenem Susceptibility in Pseudomonas aeruginosa by Hfq Rewiring of Gene Expression in Pseudomonas aeruginosa During Diauxic Growth Reveals an Indirect Regulation of the MexGHI-OpmD Efflux Pump by Hfq RNA-Seq Based Transcriptional Profiling of Pseudomonas Aeruginosa Pa14 After Short- and Long-Term Anoxic Cultivation in Synthetic Cystic Fibrosis Sputum MediumCitation
Elisabeth Sonnleitner, Petra Pusic, Michael T. Wolfinger, Udo Blasi
Front. Microbiol. 11:1001 (2020) | doi:10.3389/fmicb.2020.01001 | PDFSee Also
Marlena Rozner, Ella Nukarinen, Michael T. Wolfinger, Fabian Amman, Wolfram Weckwerth, Udo Blaesi, Elisabeth Sonnleitner
Front. Microbiol. 13:919539 (2022) | doi:10.3389/fmicb.2022.919539 | PDF
Karin Tata, Sarah G. Paltnig, Ewald H. Schwarz, Marija Mair, Michael T. Wolfinger, Elisabeth Sonnleitner, Wolfgang Schuster, Udo Blasi
PLoS ONE 11:e0147811 (2016) | doi:10.1371/journal.pone.0147811 | PDF