A conserved G-quadruplex in the Zika virus 3' terminal region
This study examines evidence for a conserved G-quadruplex in the Zika virus 3' terminal region and discusses what the observed DDX17 interaction does, and does not, imply.
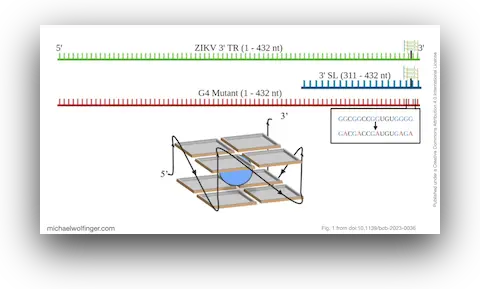
Zika virus (ZIKV) is a positive-strand flavivirus whose untranslated terminal regions contain structured RNA elements with likely regulatory roles. In this study, we focused on a G-rich segment in the 3' terminal region and asked two fairly specific questions: is there convincing evidence that this segment forms a conserved G-quadruplex, and how does that element interact with host helicases?
G-quadruplexes are structurally distinct RNA elements formed by stacked guanine quartets. In viral RNAs they are interesting because they may compete with alternative conformations, alter accessibility, or modulate how host factors engage a structured region. That does not make every predicted G4 biologically relevant, but it does make them worth testing when the underlying sequence signal is conserved.
The central result is that the candidate ZIKV G4 is not just a short-oligo artifact. Across available ZIKV isolates, the G-rich segment is conserved, and biochemical assays support G4 formation in the context of a larger 3' terminal region transcript. Pyridostatin and BG4 binding provided experimental support for that interpretation. This moves the discussion from sequence speculation to a more defensible structural observation.
The second important result is that human DEAD-box helicases interact with this region. Both DDX3X132-607 and DDX17135-555 bind the 3' terminal region, and the DDX17 construct used in the study unfolds the G4. Mechanistically, that is interesting because it suggests the local RNA ensemble is not static. Host proteins may reshape this part of the viral RNA, potentially shifting the balance between alternative conformations or affecting downstream processes that depend on terminal-region architecture.
What the study does not show, however, is just as important. It does not establish that the G4 is required for viral replication in cells, nor does it show that DDX17 binding or unwinding is directly druggable in a useful therapeutic sense. Those are hypotheses that may follow from the data, but they are not conclusions supported by the present experiments. For a system like this, the right next steps are functional perturbation, context-specific virology, and direct tests of how the structured region behaves during infection.
This distinction matters because structured viral RNAs are easy to oversell. A conserved element plus a host-factor interaction is already a meaningful result. It tells us there is a plausible structured RNA feature in the ZIKV 3' terminal region and that at least one host helicase can remodel it. That is strong mechanistic groundwork, even before any translational claims enter the picture.
For RNA virology more broadly, this is exactly where computational and biochemical analysis work well together. Comparative sequence analysis identifies conserved candidates, structural assays test whether those candidates persist in larger transcript contexts, and protein-binding experiments begin to reveal how host factors may shape the RNA landscape. That combined view is much more informative than any one layer on its own.
For teams evaluating viral RNA structure hypotheses or RNA-protein interaction models, I also offer focused technical review and advisory support through my services page. The 3’ terminal region of Zika virus RNA contains a conserved G-quadruplex and is unfolded by human DDX17Citation
Danielle L. Gemmill, Corey R. Nelson, Maulik D. Badmalia, Higor S. Pereira, Michael T. Wolfinger, and Trushar Patel
Biochem. Cell Biol. 102(1):96–105 (2024) | doi:10.1139/bcb-2023-0036 | PDF