Xinyang flavivirus and a likely tick-only orthoflavivirus clade
Xinyang flavivirus defines a basal, likely tick-only orthoflavivirus clade and expands the comparative picture of vertebrate-independent flavivirus evolution.
This paper is interesting because it changes the comparative picture of tick-borne orthoflaviviruses in two ways at once. First, it adds a new geographic data point. Xinyang flavivirus was detected in Haemaphysalis flava ticks in China and groups with Mpulungu flavivirus from Zambia and Ngoye virus from Senegal. That means the basal clade defined earlier from African ticks is not a local curiosity. It extends at least into Asia and looks increasingly like a real, ecologically coherent lineage.
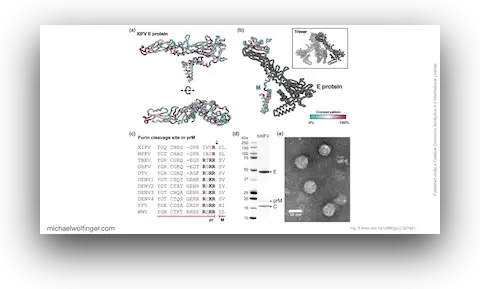
Second, the paper sharpens the argument that this lineage may be vertebrate-independent. Most tick-borne flaviviruses are discussed in terms of transmission cycles involving ticks and vertebrate hosts. XiFV, MPFV, and NGOV do not fit that expectation well. Their dinucleotide composition is closer to classical insect-specific flaviviruses than to vertebrate-infecting tick-borne flaviviruses, and XiFV shares with MPFV the absence of a furin cleavage site in prM that is otherwise common in vertebrate-associated relatives. The picture that emerges is not merely “another unusual flavivirus,” but a basal branch that may have settled on a tick-only life cycle.
That is where the structural work becomes important. The paper does not stop at phylogeny and host-range inference. It also examines the 3' UTR and shows that XiFV retains a recognizable orthoflaviviral RNA architecture, including multiple pseudoknot-containing stem-loops and a prototypical dumbbell element. In other words, even a lineage that may have abandoned vertebrate transmission still preserves a sophisticated structured non-coding region. That makes the system valuable for thinking about which RNA elements belong to the core flaviviral toolkit and which may be associated with particular host or transmission regimes.
This connects naturally to the earlier Mpulungu virus study, which first established the unusual African branch and identified distinctive xrRNA-like features in its 3' UTR. XiFV strengthens that story by showing that the lineage is broader than MPFV alone and by adding another genome against which the structural and ecological hypotheses can be tested. It also sits well beside the Zambia insect-specific flavivirus xrRNA work, where the emphasis was likewise on how non-coding RNA structure helps reveal deeper commonalities across understudied flavivirus groups.
What makes XiFV especially useful is that it sits at the intersection of discovery, comparative genomics, and evolutionary interpretation. The virus broadens the known distribution of this basal clade, supports the idea of a vertebrate-independent tick-only branch, and adds another structured 3' UTR to the small but growing set of non-coding regions that can be compared across unusual orthoflaviviruses. That combination makes the paper more than a geographic range extension. It is a clearer statement that the ecological diversity of tick-borne flaviviruses is wider than the classical vertebrate-centered model suggests.
It also fits naturally with When sequence conservation is not enough to find functional RNA structure, because XiFV reinforces how much of the informative signal sits at the level of conserved architecture rather than straightforward sequence identity. Xinyang flavivirus, from Haemaphysalis flava ticks in Henan province, China, defines a basal, likely tick-only flavivirus clade An African Tick Flavivirus Forming an Independent Clade Exhibits Unique Exoribonuclease-Resistant RNA Structures in the Genomic 3’-Untranslated RegionCitation
Lan-Lan Wang, Qia Cheng, Natalee D. Newton, Michael T. Wolfinger, Mahali S. Morgan, Andrii Slonchak, Alexander A. Khromykh, Tian-Yin Cheng, Rhys H. Parry
J. Gen. Virol. 105(5) (2024) | doi:10.1099/jgv.0.001991 | PDFSee Also
Hayato Harima, Yasuko Orba, Shiho Torii, Yongjin Qiu, Masahiro Kajihara, Yoshiki Eto, Naoya Matsuta, Bernard M. Hang’ombe, Yuki Eshita, Kentaro Uemura, Keita Matsuno, Michihito Sasaki, Kentaro Yoshii, Ryo Nakao, William W. Hall, Ayato Takada, Takashi Abe, Michael T. Wolfinger, Martin Simuunza, Hirofumi Sawa
Sci. Rep. 11:4883 (2021) | doi: 10.1038/s41598-021-84365-9 | PDF