From structure to function in Musashi-RNA complexes
This review article surveys how computational modeling, molecular dynamics, and AI-derived structures help explain Musashi-RNA recognition in both cellular regulation and viral pathogenesis.
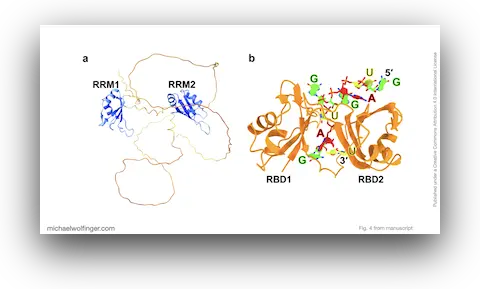
Musashi proteins sit at an interesting intersection of RNA biology, structural modeling, and disease relevance. They are sequence-selective RNA-binding proteins that regulate translation and cell-state decisions, but they have also emerged in viral contexts, most notably through proposed interactions with structured flaviviral RNAs. That combination makes Musashi a good example of a system where the mechanistic question is not just whether binding occurs, but how structural recognition translates into biological function.
This review brings together the computational work on that question. Rather than presenting a single new simulation pipeline, it synthesizes what molecular modeling, docking, molecular dynamics, and recent AI-assisted structure prediction have taught us about Musashi-RNA recognition. The focus is on how the two RNA-binding domains of Musashi engage short sequence motifs, how local RNA context modulates accessibility, and how these interactions can be interpreted in both endogenous regulatory RNAs and viral genomes.
One useful contribution of the article is that it connects several strands of prior work that are often read separately. On one side are studies of canonical Musashi recognition, where the problem is binding specificity and domain-level interaction geometry. On the other are virus-oriented analyses asking whether Musashi can plausibly recognize motifs embedded in flaviviral untranslated regions, and what that could imply for replication or neuropathology. The review argues that these are not isolated topics: the same structural principles have to explain both classes of observations.
For that reason, the paper is best read as a bridge between atomistic modeling and functional interpretation. It emphasizes that Musashi-RNA recognition depends on more than a short consensus motif in sequence space. RNA presentation, local fold, and dynamic rearrangement all influence whether a candidate site is likely to be bound in a biologically meaningful way. That is exactly where computational approaches remain useful: not as substitutes for experiment, but as tools for ranking plausible binding modes, testing structural hypotheses, and identifying which motifs deserve deeper validation.
The review also reflects a broader change in RNA structural biology. AlphaFold-class models, improved docking strategies, and longer-timescale simulation workflows have made it easier to generate mechanistic hypotheses for RNA-protein complexes, but the hard part remains interpretation. In the Musashi field, the important advance is not simply higher-confidence coordinates. It is the ability to move from static structural models toward more explicit explanations of specificity, competition, and context dependence in RNA recognition.
From my perspective, that is what makes this article worthwhile. It consolidates a line of work spanning Musashi binding to cellular RNAs, structural refinement of Musashi-RNA complexes, and possible links to viral pathogenesis. For readers interested in RNA-Protein interaction, 3D modeling, or the realistic use of AI in structural biology, it provides a compact map of the field and a clear rationale for where computation can genuinely add insight. From Structure to Function: Computational Insights into Musashi-RNA Complexes in the Context of Viral Pathogenesis and Beyond Theoretical studies on RNA recognition by Musashi 1 RNA-binding protein RNA-protein complex refinement using AI modeling and docking Musashi Binding Elements in Zika and Related Flavivirus 3’UTRs: A Comparative Study in SilicoCitation
Nitchakan Darai, Leonhard Sidl, Thanyada Rungrotmongkol, Peter Wolschann, Michael T. Wolfinger
Sci. Asia 51S(1) 2025s013:1-10 (2025) | doi:10.2306/scienceasia1513-1874.2025.s013 | PDFSee Also
Nitchakan Darai, Panupong Mahalapbutr, Peter Wolschann, Vannajan Sanghiran Lee, Michael T. Wolfinger, Thanyada Rungrotmongkol
Sci. Rep. 12:12137 (2022) | doi:10.1038/s41598-022-16252-w | PDF
Nitchakan Darai, Kowit Hengphasatporn, Peter Wolschann, Michael T. Wolfinger, Yasuteru Shigeta, Thanyada Rungrotmongkol, Ryuhei Harada
B. Chem. Soc. Jpn. 96(7):677-685 (2023) | doi:10.1246/bcsj.20230092 | PDF
Adriano de Bernardi Schneider, Michael T. Wolfinger
Sci. Rep. 9(1):6911 (2019) | doi:10.1038/s41598-019-43390-5 | PDF