Bayesian approximation of RNA folding times
This workshop paper introduces the core KinPFN idea: approximating RNA first-passage-time distributions with a prior-data fitted network trained on synthetic folding-time priors.
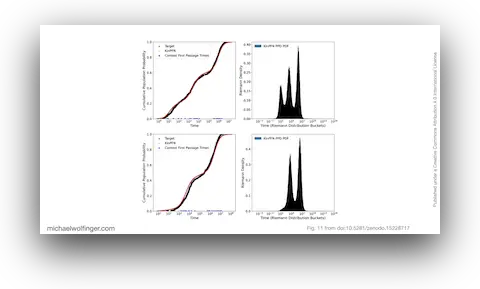
This workshop paper presents the core idea behind KinPFN in a compact form: model RNA folding-time distributions directly, instead of recomputing them through thousands of simulator runs for every new case. The target quantity is the cumulative distribution of first passage times, a useful kinetic summary because it reflects how quickly different fractions of molecules reach a structure of interest.
The proposed solution uses prior-data fitted networks, trained not on a large corpus of experimentally measured RNAs, but on synthetic multi-modal distributions chosen to resemble the behavior of real folding-time data. Given just a few example folding times from a kinetics simulator, the model predicts the posterior distribution of folding times and from that the full cumulative distribution function. That makes the method especially attractive where only a small kinetic sample is available but a full distribution estimate would be useful.
What makes this interesting for RNA folding kinetics is the combination of speed and modularity. The method does not require rewriting the underlying simulator or abandoning physically motivated kinetic models. Instead, it acts as an approximation layer that reduces the number of expensive simulations needed for downstream interpretation. The workshop paper therefore functions as a proof of principle for integrating modern probabilistic machine learning into RNA kinetics workflows.
Relative to the full conference paper, this version is shorter and more focused on the central idea, but it already makes the key argument clearly: approximating folding-time distributions can be enough for many practical tasks, and those approximations can be learned efficiently from a synthetic prior. For anyone interested in the intersection of RNA folding kinetics and AI, this paper is a useful entry point into the broader KinPFN project.
It also connects directly to Why kinetic folding matters in RNA design, because many design decisions do not require a perfect kinetic simulation. They require a fast and credible way to compare whether one candidate is likely to behave more cleanly than another. Bayesian Approximation of RNA Folding TimesCitation
Dominik Scheuer, Frederic Runge, Jörg K.H. Franke, Michael T. Wolfinger, Christoph Flamm, Frank Hutter
ICLR 2025 Workshop on AI for Nucleic Acids (2025) | doi:10.5281/zenodo.15228717 | PDF