KinPFN for RNA folding kinetics
KinPFN uses prior-data fitted networks to approximate first-passage-time distributions for RNA folding kinetics orders of magnitude faster than direct simulation.
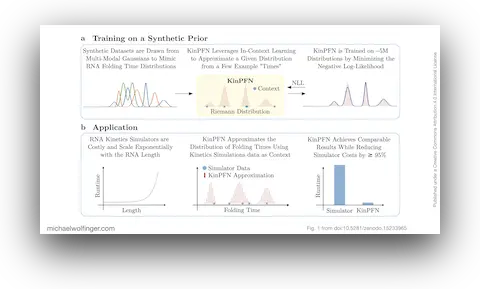
RNA folding kinetics is often summarized in terms of first passage times: how long it takes a molecule to reach a target structure for the first time. Those distributions are informative because they capture more than a single mean folding time, but obtaining them requires many stochastic simulations and quickly becomes expensive. This paper asks whether the entire folding-time distribution can be approximated directly from a small number of example simulations.
The proposed answer is KinPFN, a prior-data fitted network trained on synthetic multi-modal distributions that mimic the shape of RNA first-passage-time distributions. Instead of learning from large collections of labeled RNA molecules, the model is trained to do approximate Bayesian inference on generic folding-time distributions. At application time, it receives only a few observed folding times as context and predicts the full cumulative distribution function in a single forward pass.
That framing is important. The method is not replacing physical simulators with a black box trained on a fixed benchmark set. It is designed as a fast probabilistic approximation layer that can sit on top of existing kinetics tools such as Kinfold and return a useful estimate of the folding-time distribution long before exhaustive simulation would finish. In practice, the paper reports speed-ups of at least 95% while preserving the overall shape of the distribution sufficiently well for downstream analysis.
The real value of the approach is that it makes distribution-level reasoning about folding kinetics much more accessible. Instead of reducing kinetics to a single summary statistic, one can compare broad versus narrow first-passage distributions, detect inefficient folding processes, and screen many more candidates than direct simulation alone would allow. The paper illustrates this in analyses of eukaryotic RNAs and folding-efficiency case studies, where the quality of the approximation is already good enough to be practically informative.
For RNA folding kinetics this is a useful conceptual shift. It shows that machine learning can accelerate kinetic analysis without having to learn RNA folding from scratch. By focusing on posterior approximation rather than direct structure prediction, KinPFN becomes a practical tool for kinetic RNA design workflows, where many candidate sequences need to be compared quickly but full simulation remains computationally restrictive.
That is also why it fits naturally with Why kinetic folding matters in RNA design. In both cases, the practical value comes from deciding which candidates deserve closer attention before a project becomes expensive. KinPFN: Bayesian Approximation of RNA Folding Kinetics using Prior-Data Fitted NetworksCitation
Dominik Scheuer, Frederic Runge, Jörg K.H. Franke, Michael T. Wolfinger, Christoph Flamm, Frank Hutter
The Thirteenth International Conference on Learning Representations (ICLR'25) (2025) | doi:10.5281/zenodo.15233965 | PDF