Rational design of mechanically active RNAs
This Nucleic Acids Research paper shows that synthetic xrRNAs can be designed from topological rules and validated experimentally as mechanically active RNAs.
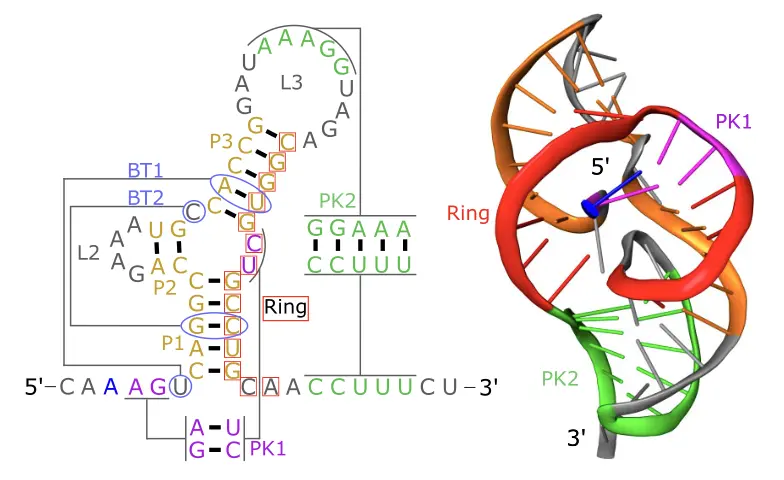
This paper extends the xrRNA story from comparative RNA virology into explicit RNA engineering. Exoribonuclease-resistant RNAs are structured viral elements that stall 5' to 3' decay through a threaded, mechanically resistant fold. The central question addressed here is whether that function can be specified de novo, rather than inherited from a natural sequence that already encodes it.
The work approaches topology as part of the design objective. Structural features relevant for XRN1 resistance are reduced to an explicit symbolic model, which is then used to generate synthetic candidates that are filtered computationally and validated experimentally. The resulting workflow combines structural constraints, ensemble-based screening, three-dimensional modelling, and molecular-dynamics selection for ring closure and directional force resistance.
One of the clearest mechanistic observations concerns the unequal contribution of the two pseudoknots to xrRNA stability. Pseudoknot 2 acts as the decisive gatekeeper of mechanical resistance, whereas pseudoknot 1 contributes additional stabilization once the fold is established. This distinction turns what initially appears as a descriptive structural feature into a tractable design rule.
Three synthetic constructs were used to progressively narrow the design space. syn-xrRNA1 approaches the intended topology but remains comparatively weak in vitro. syn-xrRNA2 improves the geometry of the critical pseudoknot region and reaches wild-type-like XRN1 resistance. syn-xrRNA3 goes a step further: much of the recognizable sequence signal is removed, yet the RNA still folds into a functional threaded architecture and efficiently resists XRN1 degradation.
To my knowledge, this is the first example of a fully de novo designed RNA in which a mechanically active fold of this complexity was specified from topological constraints and then validated experimentally. In that sense, the paper marks a genuine shift in how RNA design can be approached. Mechanically active RNA elements become accessible through topology and geometry rather than through direct sequence imitation of natural viral RNAs.
For synthetic biology and therapeutic RNA engineering, this opens the possibility of tuning decay resistance and transcript stability without importing long native viral sequence segments wholesale. At the same time, the work remains fundamentally a study in RNA architecture. The main result is that mechanical function can be preserved even after evolutionary sequence ancestry has effectively been removed.
The broader implications extend well beyond flavivirus RNA biology. If mechanically active RNA elements can be specified de novo, then decay control becomes an engineering variable rather than a borrowed viral feature. That changes how one can think about stabilizing synthetic transcripts, shaping RNA lifetime in therapeutic settings, or introducing programmable decay barriers into larger regulatory designs. In RNA therapeutics, where persistence, dosage, and degradation behaviour are often as important as coding capacity, this kind of design principle could become a useful addition to the current toolbox. More generally, the work shows that higher-order RNA architecture itself can be treated as a design substrate, not only as something discovered retrospectively in natural molecules.
The present study also closes a conceptual loop with earlier work on natural xrRNAs and structured viral RNAs. Comparative analysis originally established the relevant folds in nature. The current paper asks which parts of that function survive once sequence history is removed and only the underlying topological logic is retained.
The January preprint-stage note remains online as a shorter record of how the project was framed before the peer-review process. Rational design of mechanically active RNAs: de novo engineering of functional exoribonuclease-resistant RNAsCitation
Jule Walter, Leonhard Sidl, Katrin Gutenbrunner, Denis Skibinski, Tim Kolberg, Ivo L. Hofacker, Hua-Ting Yao, Mario Mörl, Michael T. Wolfinger
Nucleic Acids Res. 54(9):gkag473 (2026) | doi:10.1093/nar/gkag473 | Article | PDF