mRNA degradation occurs on ribosome complexes in Drosophila cells
This study shows that bulk and miRNA-guided mRNA degradation in Drosophila cells occurs on ribosome-associated messenger ribonucleoprotein complexes, linking decay machinery, translation, and high-throughput sequencing of decapped intermediates.
This paper sits in a different part of my publication list than the RNA structure and folding work, but in hindsight it connects to it more than it may seem at first glance. Before I started to investigate viral xrRNAs and the ways in which structured RNAs can resist exonucleolytic decay, I was interested in a more general question: where does 5' to 3' mRNA degradation actually take place in the cell, and how tightly is it coupled to translation?
At the time, the coupling between translation and mRNA turnover was already widely appreciated, but the physical location of that coupling was still not entirely clear. One could imagine a handoff model in which translating ribosomes and decay machineries act in sequence, or a more direct model in which degrading mRNAs remain associated with ribosome-containing complexes during turnover. This study addressed that distinction experimentally in Drosophila cells and asked whether the same picture also applies to miRNA-mediated decay.
The approach combined several complementary readouts. Polysome profiling and ribosome affinity purification were used to isolate ribosome-associated complexes. These preparations were then probed for canonical translation factors, general deadenylation and decapping components, and key miRNA pathway proteins such as AGO1 and GW182. The crucial control was that these associations depended on intact RNA, arguing that the decay factors were not simply sticking nonspecifically to ribosomal proteins but were traveling with messenger ribonucleoprotein complexes engaged with mRNA.
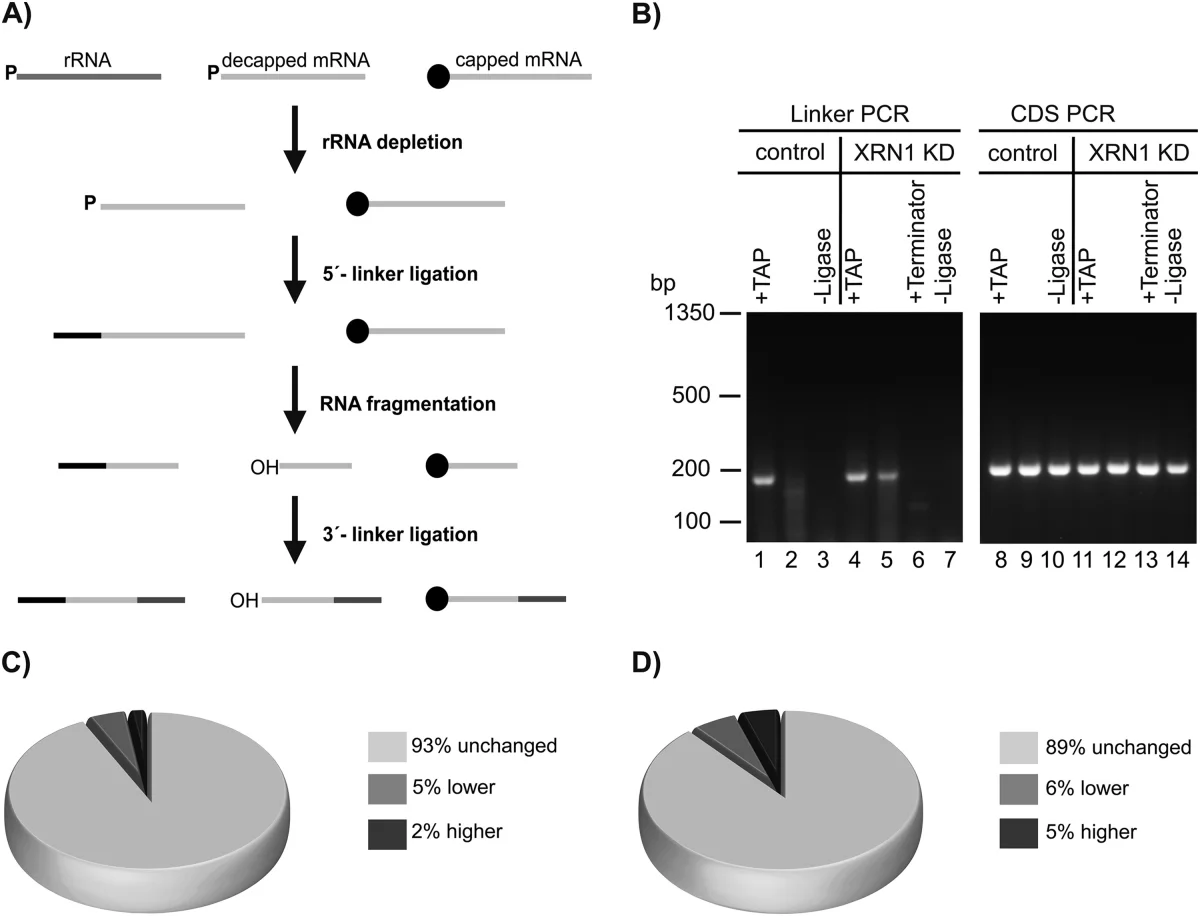
The most informative part of the paper, in my view, is the sequencing-based analysis of decapped degradation intermediates. The workflow captured 5' monophosphorylated mRNA fragments from ribosome complexes and compared them with corresponding fragments from whole-cell lysate. That moved the study beyond a few marker transcripts or individual immunoblots. Instead, it asked the question transcriptome-wide: are decapped intermediates generally depleted from ribosome-associated fractions, or do they remain there at similar relative abundance?
The answer was strikingly broad. Roughly 93% of the detected decapped fragments were present at essentially the same relative abundance in ribosome complexes as in the total lysate. That is a strong argument against the idea that decapping is followed by rapid physical separation from translational assemblies for most transcripts. Rather, the data support a model in which bulk 5' to 3' decay commonly proceeds on ribosome-associated mRNPs, with miRNA-mediated decay fitting into the same overall framework rather than constituting a fully separate compartmentalized process.
That result matters biologically because it shifts the emphasis from static cellular compartments to dynamic functional coupling. It suggests that translation, deadenylation, decapping, and exonucleolytic degradation should often be understood as parts of the same continuum of messenger RNA handling. For miRNA biology this was especially relevant: AGO1 and GW182 were not just abstract regulatory labels but factors connected to ribosome-associated decay states of target RNAs.
Seen from later work on xrRNAs, this paper also sharpened the next question. If much of 5' to 3' decay is happening on ribosome-associated messenger complexes, then structured RNA elements that can impede exonucleases become even more interesting, because they do not act in an isolated degradation chamber. They act in the middle of an already crowded and highly coordinated post-transcriptional environment. In that sense, this study predates the xrRNA projects but helped define the decay-centered perspective from which those later questions became compelling.
So although this is not an RNA structure paper, it is an important mechanistic one. It combines classical biochemical fractionation with a transcriptome-wide sequencing readout to make a fairly clean point: in Drosophila cells, both general and miRNA-mediated mRNA degradation are closely tied to ribosome-associated complexes. For anyone interested in how gene expression is controlled after transcription, that is a useful result to keep in mind. General and miRNA-mediated mRNA degradation occurs on ribosome complexes in Drosophila cellsCitation
Sanja Antic, Michael T. Wolfinger, Anna Skucha, Stefanie Hosiner and Silke Dorner
Mol. Cell. Biol. 35(13), 2309-2320 (2015) | doi:10.1128/MCB.01346-14 | PDF