Strukturierte RNAs in Viren (in German)
In this mini-review we discuss the concept of RNA structure conservation in viruses, using exoribonuclease-resistant RNAs from flaviviruses as prominent examples
This article is a short review, but it covers a central idea that has become increasingly important across virology: many viral genomes do not just encode proteins, they also encode structured RNA elements that actively control the infection cycle. In flaviviruses, the 3'UTR is a particularly rich example because it contains conserved folds that regulate replication, translation, host adaptation, and immune evasion.
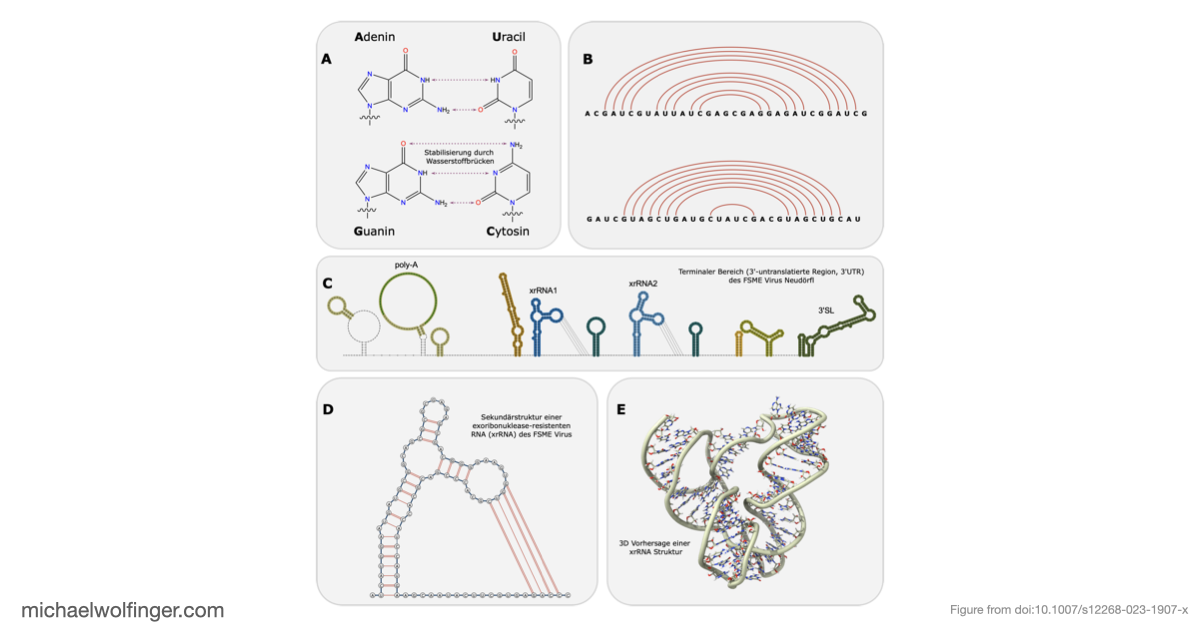
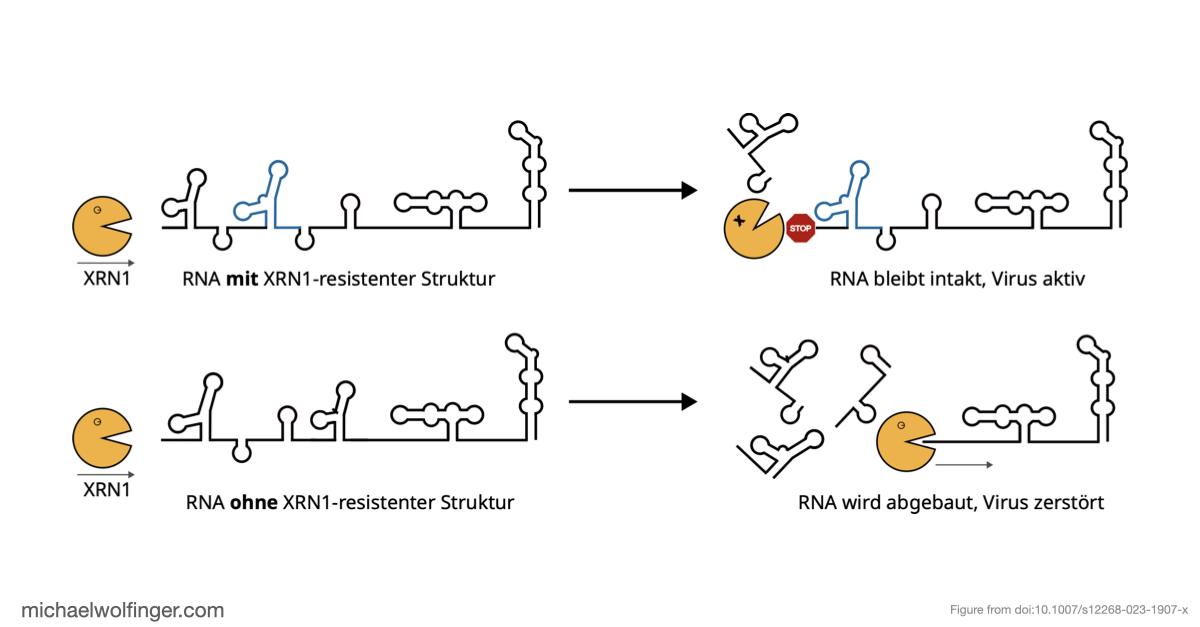
The focus here is on exoribonuclease-resistant RNAs, or xrRNAs. These elements are interesting because they do not merely bind a host factor or present a passive recognition site. They act mechanically. Their three-dimensional fold stalls the cellular XRN1 exoribonuclease and thereby generates subgenomic flaviviral RNAs (sfRNAs). Those sfRNAs are now known to contribute to pathogenicity, host-range effects, and immune modulation in multiple flavivirus groups.
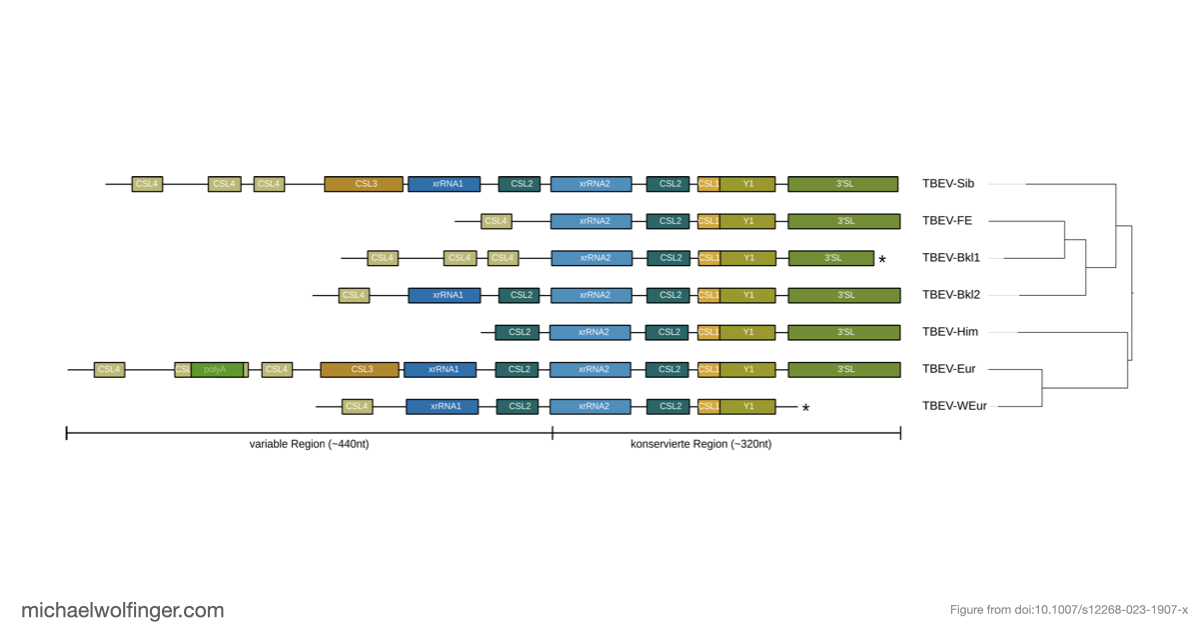
What makes xrRNAs especially attractive from a structural-biology perspective is that they illustrate RNA function in a very direct way. Their activity depends on conserved topology rather than simple sequence identity. That means they are an ideal example of why RNA structure conservation matters in virology: two viral RNAs can diverge substantially at sequence level while preserving a fold that continues to perform the same mechanistic task.
The review therefore works on two levels. At the biological level, it summarizes why structured 3'UTR elements matter for the viral life cycle and how xrRNAs help viruses manipulate host RNA decay pathways. At the conceptual level, it argues that RNA structures should be treated as evolvable functional modules, not just decorative features of untranslated regions. That framing connects this short piece to a much broader body of comparative work on conserved viral RNAs.
Another useful aspect of the article is that it points beyond natural virus biology toward design. If an RNA fold can reliably block an exoribonuclease in nature, then the same principle can potentially be repurposed in synthetic systems. That is where the connection to RNA design and future therapeutic applications comes in. xrRNAs are interesting not only because they explain flavivirus biology, but because they may serve as modular regulatory parts in engineered RNAs.
This is also why the topic has become relevant in the context of RNA therapeutics and mRNA technology. Structured RNA elements that control stability or degradation are obvious candidates for engineering transcript lifetime and behavior. The point is not that viral xrRNAs can simply be copied into therapeutic constructs without modification, but that they provide a proven natural design principle for mechanically active RNA control.
The piece functions as a compact bridge between comparative viral RNA structure analysis, xrRNA discovery, and later efforts in synthetic and mechanically active RNA design. Even though the article itself is shorter and more magazine-style than a primary research paper, the underlying concept is foundational and deserves more context than a brief teaser. Readers who want to follow that thread into primary research can move from here to Discoveries of Exoribonuclease-Resistant Structures of Insect-Specific Flaviviruses Isolated in Zambia or to Rational design of mechanically active RNAs.
For a more general discussion of why sequence similarity alone often fails to capture these conserved viral RNA elements, see When sequence conservation is not enough to find functional RNA structure. Strukturierte RNAs in Viren (in German)Figures and Data
Citation
Roman Ochsenreiter, Michael T. Wolfinger
Biospektrum 29(2):156-158 (2023) | | doi:10.1007/s12268-023-1907-x | PDF