Co-transcriptional riboswitch modeling with ViennaRNA
A landscape-based method for modeling how cotranscriptional folding and ligand binding interact in kinetically controlled riboswitches, illustrated with the 2'dG riboswitch from Mesoplasma florum.
Riboswitches are a good reminder that RNA function is often kinetic rather than purely thermodynamic. For many ligand-sensing RNAs, the biologically relevant question is not simply which structure has the lowest free energy at equilibrium. What matters is which structures become available during transcription, how long they persist, and whether ligand binding happens early enough to redirect the folding pathway before an alternative conformation takes over.
That is the problem addressed in this paper. The goal is to model cotranscriptional RNA-ligand interaction dynamics efficiently enough to say something mechanistic about a kinetically controlled riboswitch, without having to rely entirely on huge numbers of direct folding trajectories. The approach builds on the energy-landscape framework developed in earlier work and extends it to a setting where both the RNA chain and the ligand-binding state change over time.
The methodological core is an extension of the BARRIERS/BarMap style of coarse graining. For each transcript length, the folding landscape is partitioned into macrostates, representing basins of attraction around locally stable structures. Those macrostates are then linked between successive transcript lengths so that population densities can be transferred forward as transcription proceeds. What makes this paper new is that ligand competence is incorporated directly into that framework. Structures that can bind ligand are treated differently from those that cannot, and ligand binding or unbinding is modeled in a concentration-dependent way rather than as a static annotation.
The simplifying assumption is deliberately clear: ligand binding is treated in an all-or-none fashion for a defined binding-competent structure. That sounds restrictive, but it is exactly the kind of abstraction that makes this model useful. It keeps the state space manageable while still capturing the central competition between RNA folding timescales and metabolite binding timescales. In practice, that is the core of many riboswitch mechanisms.
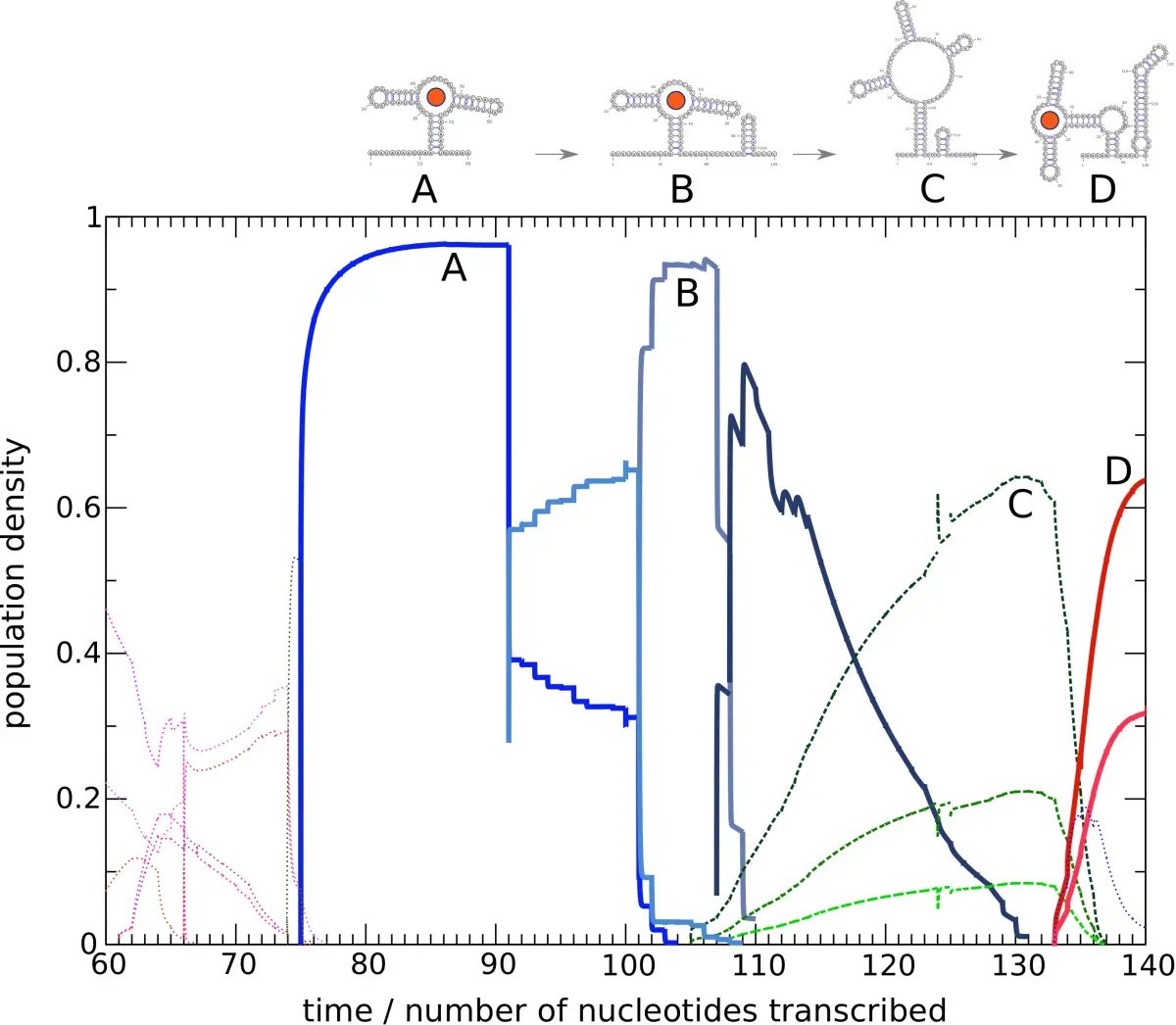
The case study is the I-A 2'-deoxyguanosine (2'dG) riboswitch from Mesoplasma florum, which had already been characterized experimentally by NMR. That makes it a strong benchmark system, because the computational model can be compared against an independently studied folding mechanism rather than against another computational prediction. The figure above captures the main idea well: as the transcript elongates, the dominant population shifts through a series of metastable states, and ligand binding changes which path remains accessible.
The main result is that this landscape-based treatment reproduces the logic of a kinetically controlled riboswitch surprisingly well. The model shows how an early binding-competent aptamer state can capture ligand and thereby steer the RNA away from competing conformations that would otherwise dominate later. The riboswitch cannot be understood from a single end-state structure alone. Its behavior depends on the timing of transcription, the accessibility of intermediate states, and the concentration-dependent opportunity for ligand capture during folding.
I still think this is one of the more useful methodological papers in the riboswitch area. It turns a vague statement like "cotranscriptional effects matter" into a concrete computational workflow. Instead of treating kinetics as an afterthought, the paper makes the dynamic landscape itself the object of study. This is relevant not only for natural riboswitches, but also for synthetic biology, where one often wants to know whether a designed switch is merely structurally plausible or actually kinetically workable.
The paper also connects two strands of RNA research that are often discussed separately: landscape-based folding kinetics and ligand-regulated RNA control. Bringing them together makes it possible to use coarse-grained kinetics not just for descriptive folding studies, but for mechanism-aware analysis of regulatory RNAs. That is a meaningful step beyond equilibrium folding and a useful basis for later in silico screening of switch designs before experimental validation.
That screening logic sits very close to Why kinetic folding matters in RNA design. A construct can look plausible at equilibrium and still fail once timing and pathway dependence are taken seriously. Efficient Computation of Cotranscriptional RNA-Ligand Interaction Dynamics NMR Structural Profiling of Transcriptional Intermediates Reveals Riboswitch Regulation by Metastable RNA ConformationsCitation
Michael T. Wolfinger, Christoph Flamm, Ivo L. Hofacker
Methods 143:70–76 (2018) | doi: 10.1016/j.ymeth.2018.04.036 | PDFSee Also
Christina Helmling, Anna Wacker, Michael T. Wolfinger, Ivo L. Hofacker, Martin Hengsbach, Boris Fürtig, Harald Schwalbe
J. Am. Chem. Soc. 139 (7):2647–56 (2017) | doi:10.1021/jacs.6b10429