Comparative genomics of flavivirus 3' UTR RNA structures
A comparative genomics analysis of flavivirus 3' UTRs that identifies conserved exoribonuclease-resistant RNAs and lineage-specific architectural variation across tick-borne, insect-specific, and no-known-vector flaviviruses.
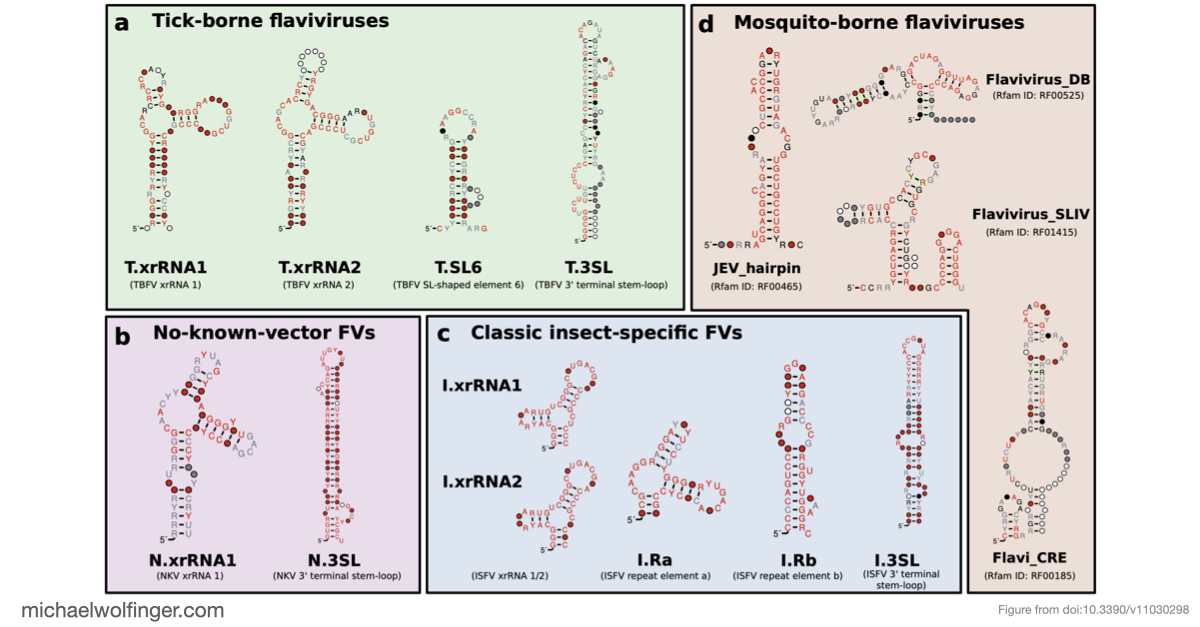
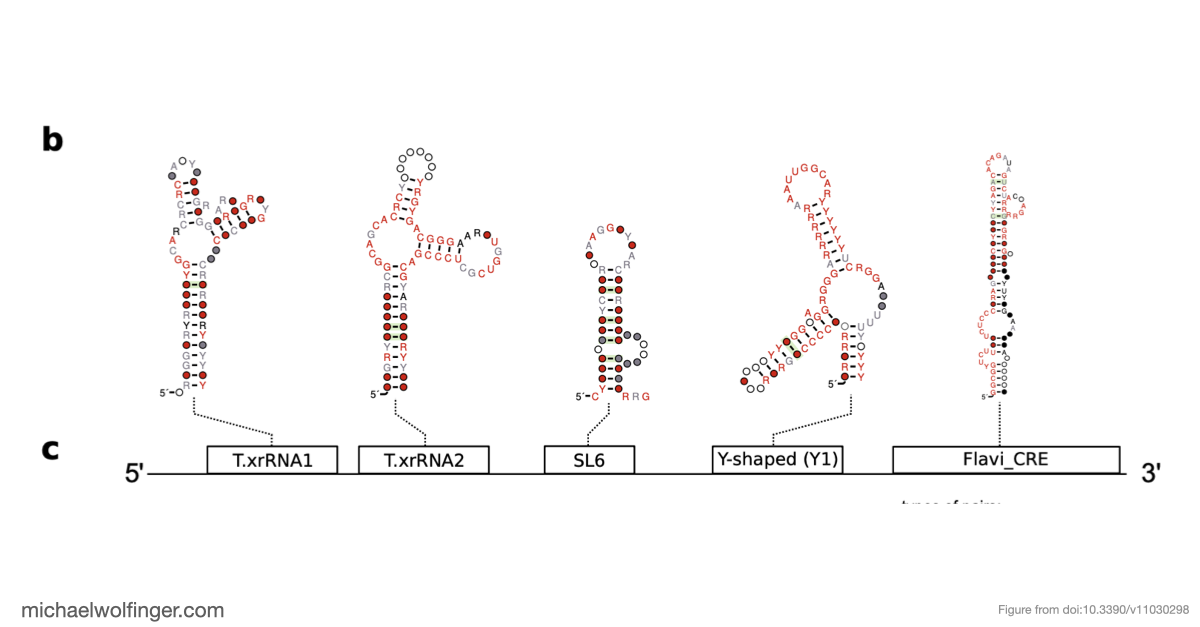
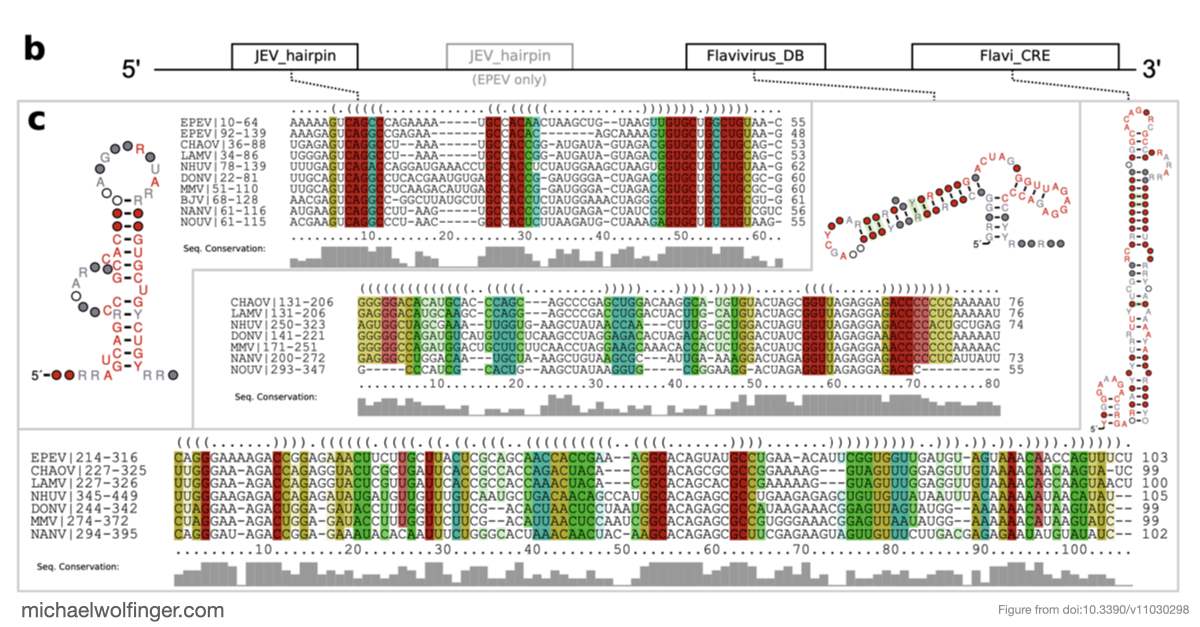
The 3' untranslated regions of flaviviruses contain far more than generic sequence context. They carry structured RNA elements involved in cyclization, replication, encapsidation, and resistance to host exonucleases. That makes them a natural target for comparative analysis, especially in viral groups where functional annotation is still sparse.
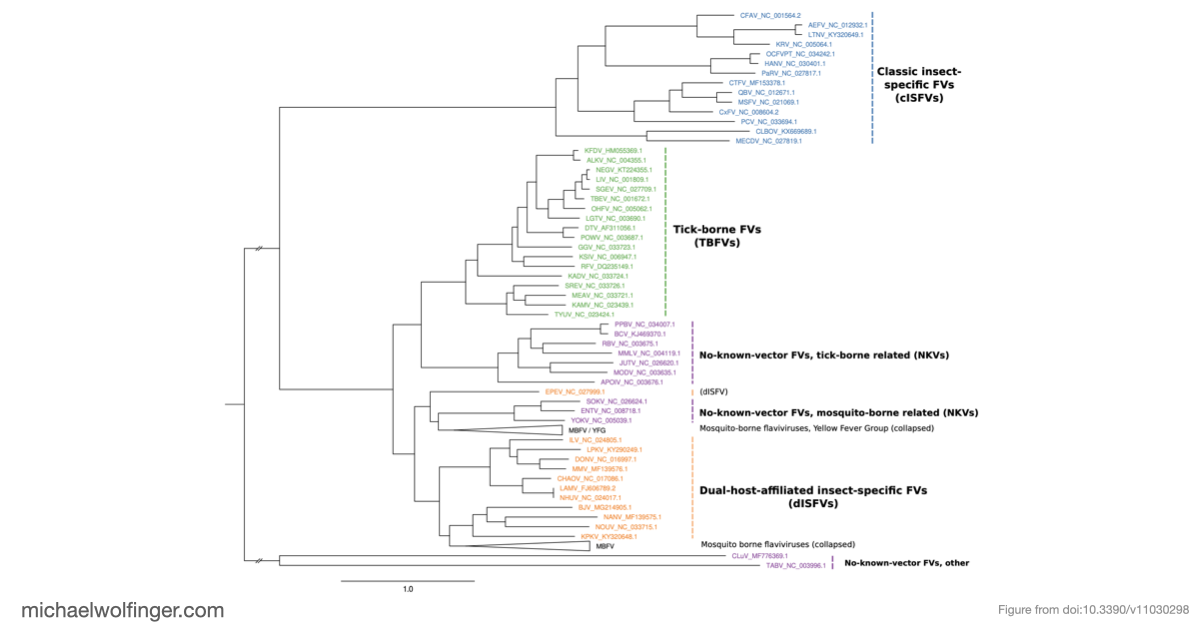
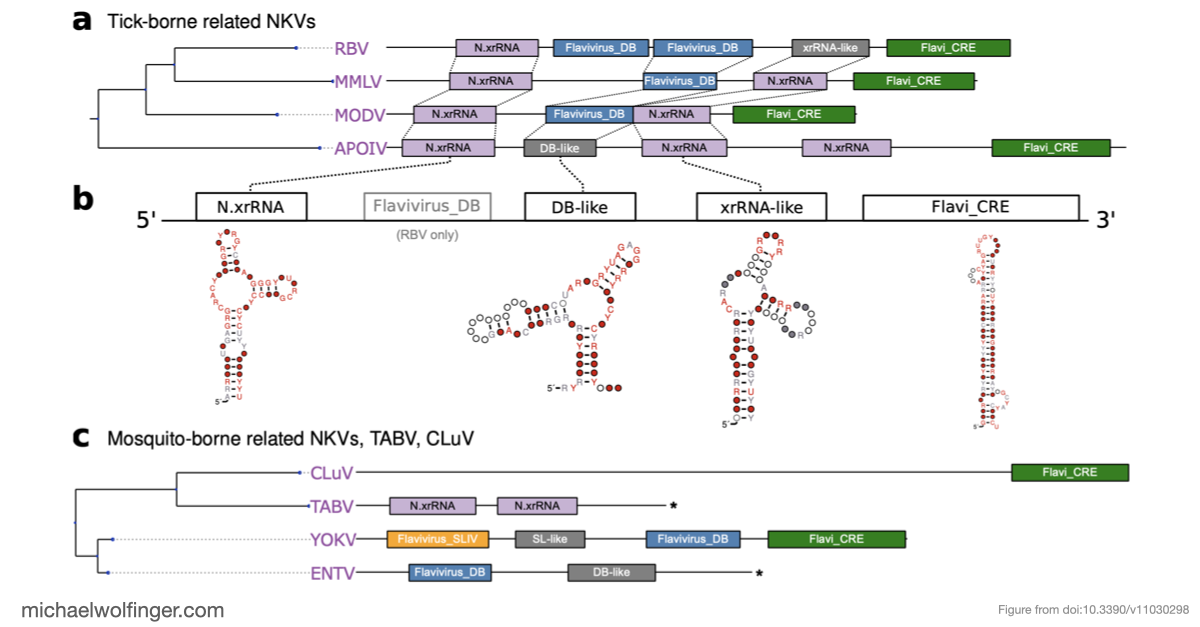
This paper takes a comparative genomics approach to the 3' UTRs of tick-borne, insect-specific, and no-known-vector flaviviruses. The question is not simply whether these viruses contain structured RNA, but how conserved architectures are distributed across lineages and where different flavivirus groups seem to have arrived at different structural solutions.
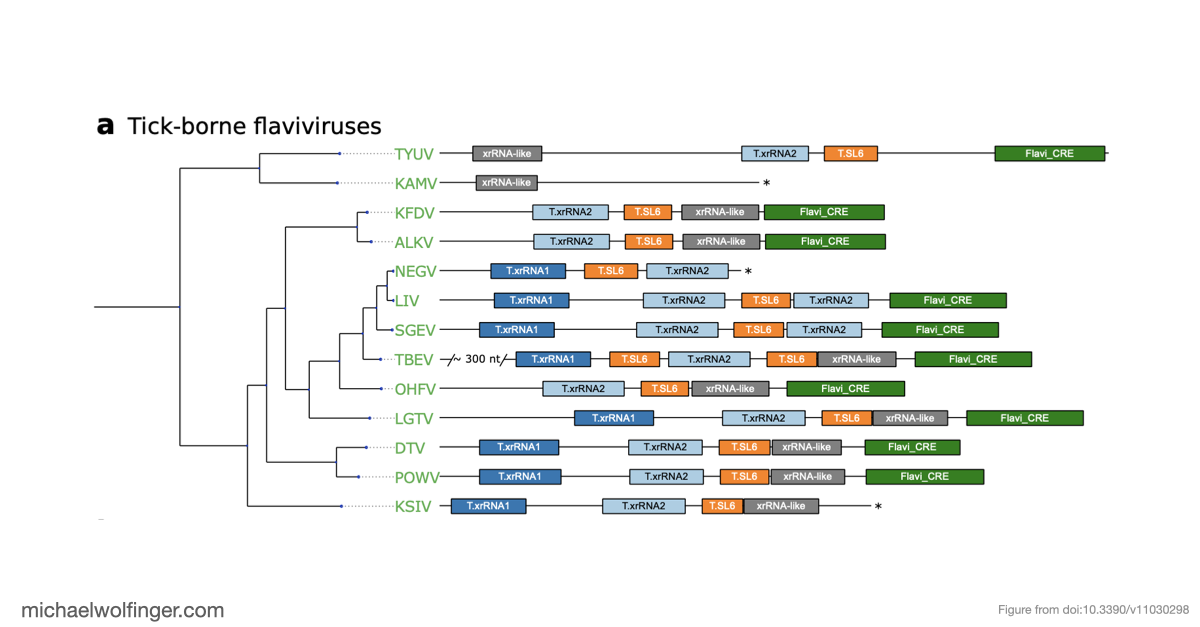
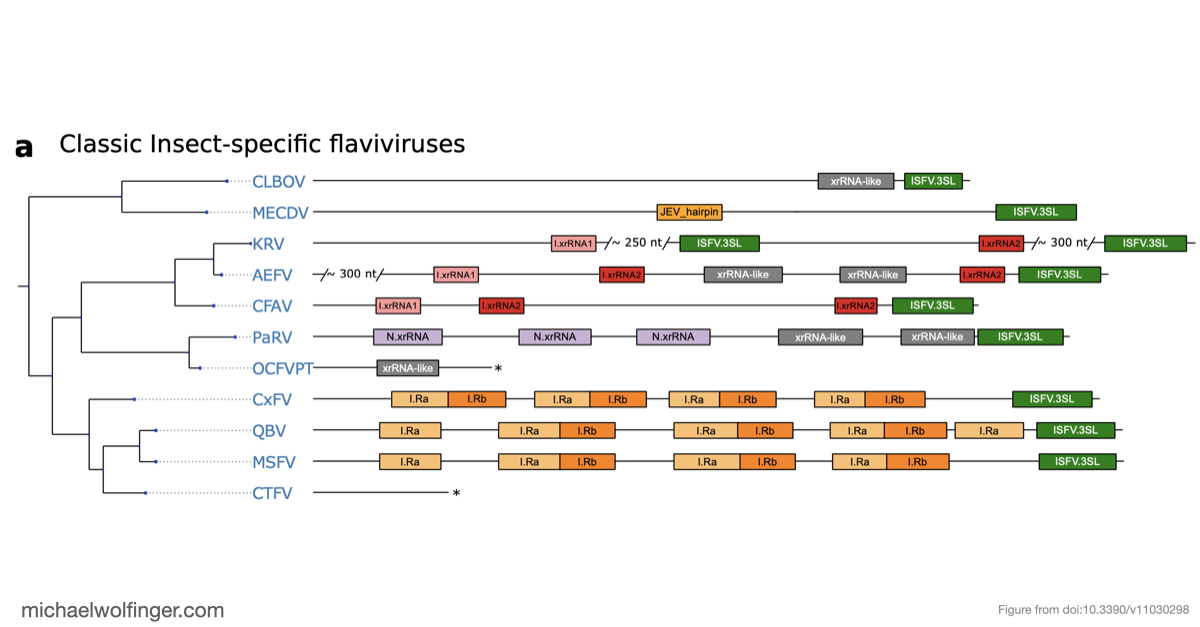
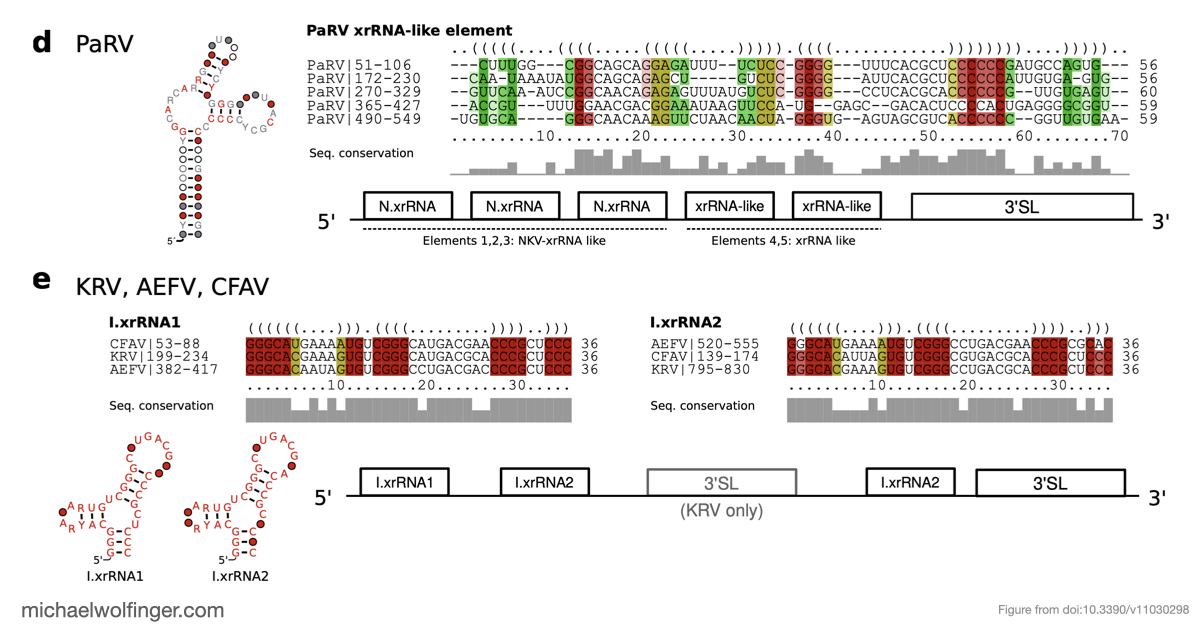
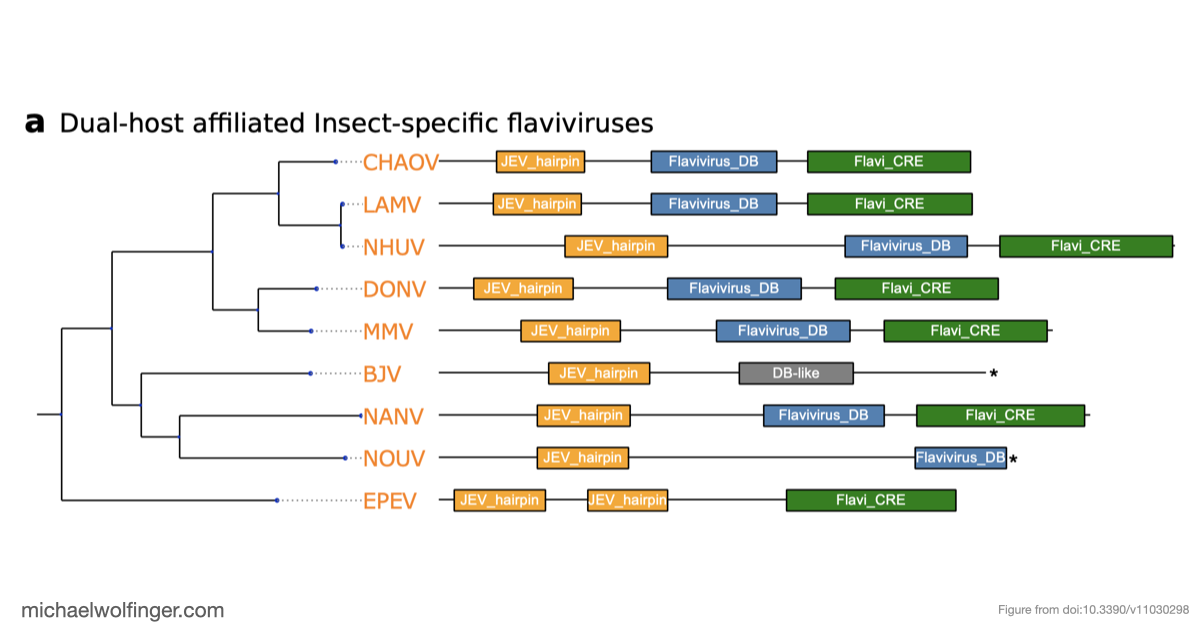
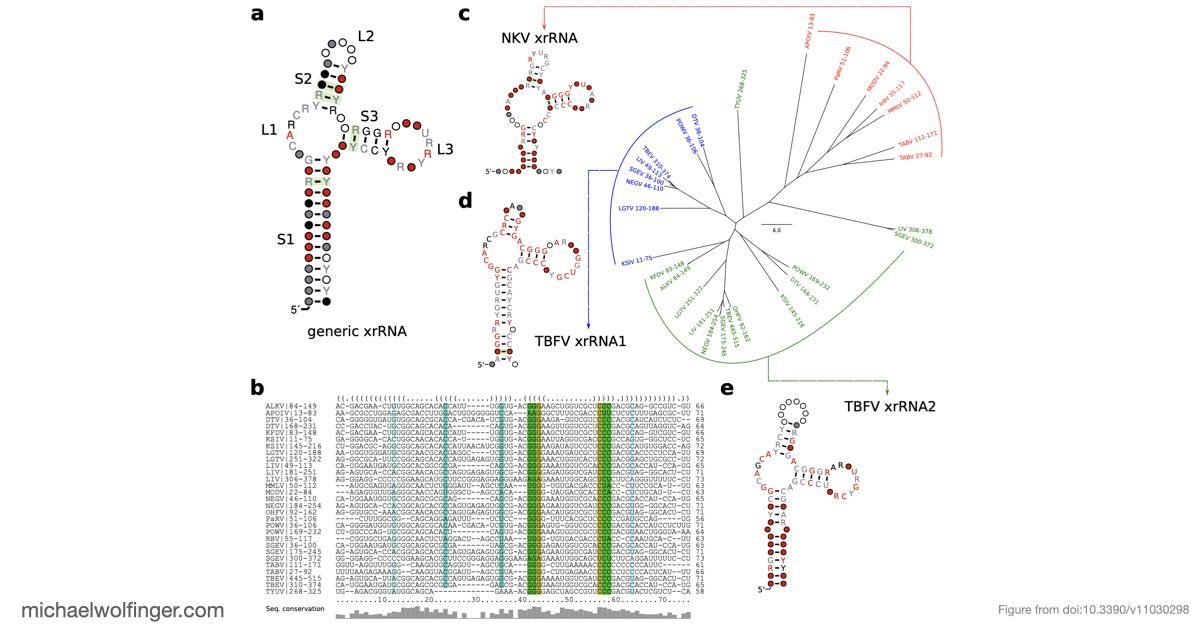
One of the main results is that exoribonuclease-resistant RNAs (xrRNAs), which had already been characterized experimentally in other flavivirus contexts, are more widely distributed than a narrow lineage-specific view would suggest. The analysis supports their presence in tick-borne and no-known-vector flaviviruses, reinforcing the idea that resistance to 5' to 3' degradation is a broadly reused structural strategy in flavivirus evolution.
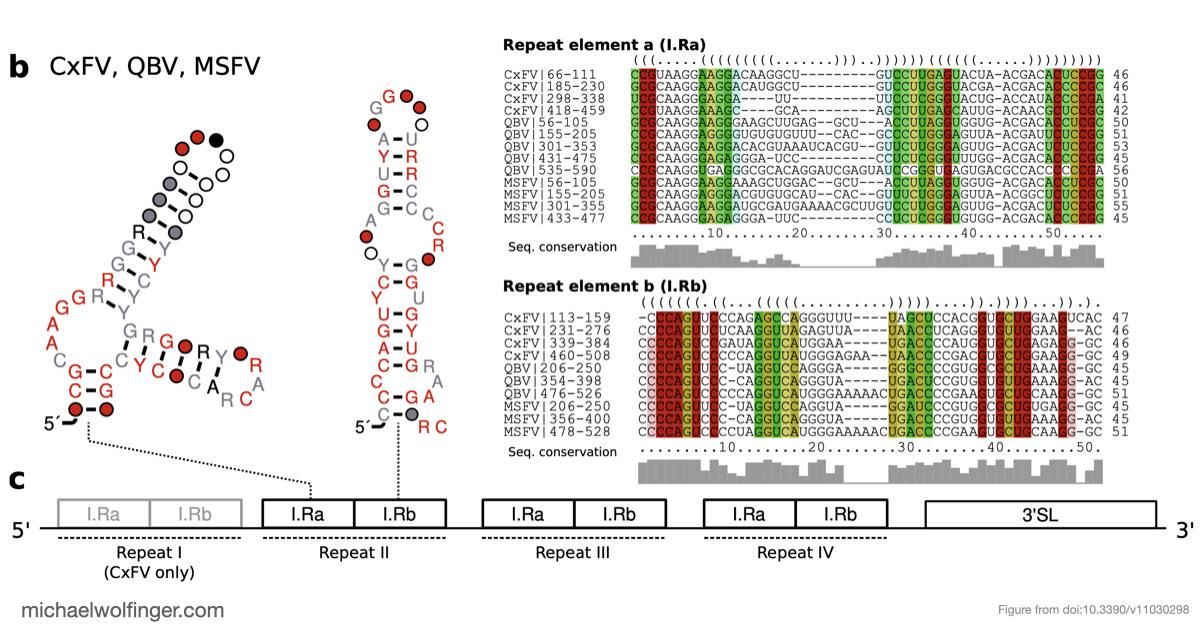
The study also points to a repeated or cascaded organization of duplicated RNA structures in insect-specific flaviviruses. That observation matters because it suggests that flavivirus 3' UTRs are not static collections of conserved motifs. They are better viewed as flexible scaffolds on which evolution can duplicate, repurpose, and elaborate structured RNA elements while maintaining core functions.
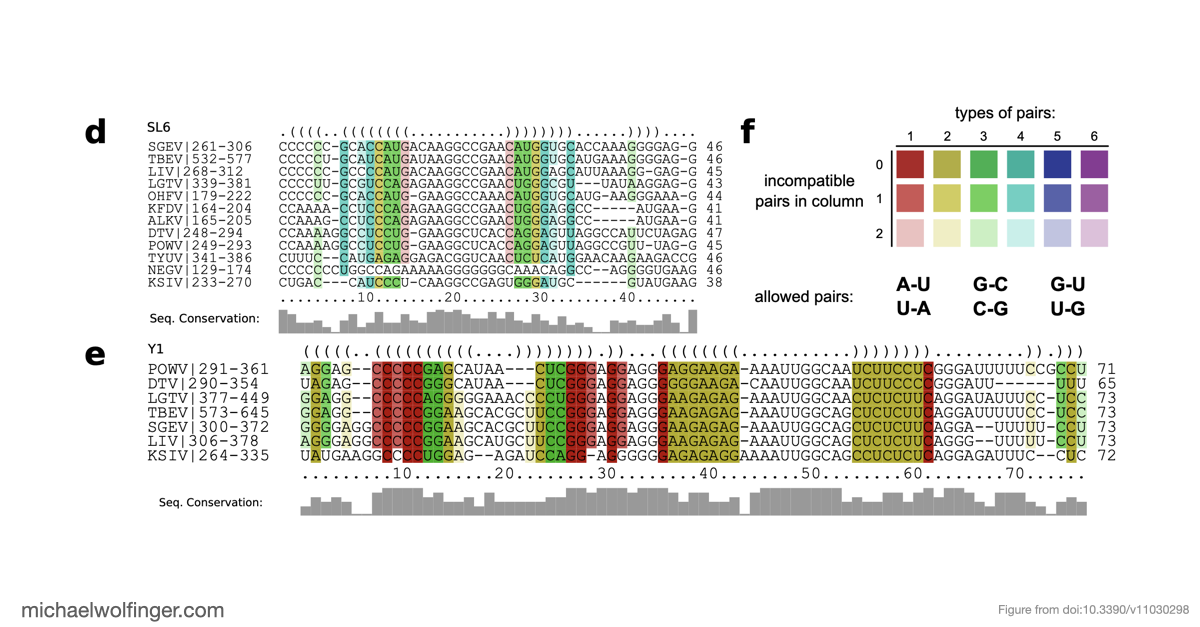
Comparative genomics is especially valuable for this kind of result. In many cases, experimental structure determination across an entire viral clade is unrealistic. Sequence comparison, consensus folding, and covariation-aware reasoning make it possible to identify plausible conserved elements first and then ask which of them are most worth testing in the lab.
For work on xrRNAs, this broader evolutionary view is particularly useful. It helps separate features that are likely to be deeply conserved from those that may have arisen independently or been remodeled in particular viral groups. In turn, that sharpens how we think about structure-function relationships in flavivirus non-coding regions.
I return to that general issue in When sequence conservation is not enough to find functional RNA structure, which argues that many of the most informative signals in viral untranslated regions live at the level of conserved architecture rather than primary sequence identity. Functional RNA Structures in the 3’UTR of Tick-Borne, Insect-Specific and No Known Vector Flaviviruses Functional RNA Structures in the 3’UTR of Mosquito-Borne Flaviviruses Evolutionary traits of Tick-borne encephalitis virus: Pervasive non-coding RNA structure conservation and molecular epidemiologyFigures and Data
Citation
Roman Ochsenreiter, Ivo L. Hofacker, Michael T. Wolfinger
Viruses 11:298 (2019) | doi:10.3390/v11030298 | PDF | FiguresSee Also
Michael T. Wolfinger, Roman Ochsenreiter, Ivo L. Hofacker
In Virus Bioinformatics, edited by Dmitrij Frishman and Manja Marz, pp65–100. Chapman and Hall/CRC Press (2021) | doi:10.1201/9781003097679-5 | Preprint PDF | Figures
Lena S. Kutschera, Michael T. Wolfinger
Virus Evol. (8):1 veac051 (2022) | doi:10.1093/ve/veac051 | PDF | Figures