Functional RNA structures in the 3'UTR of Mosquito-Borne Flaviviruses
This chapter compares conserved functional RNA elements in the 3' UTRs of mosquito-borne flaviviruses, including xrRNAs, dumbbells, and terminal stem-loops.
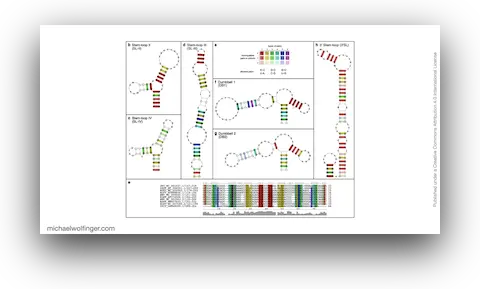
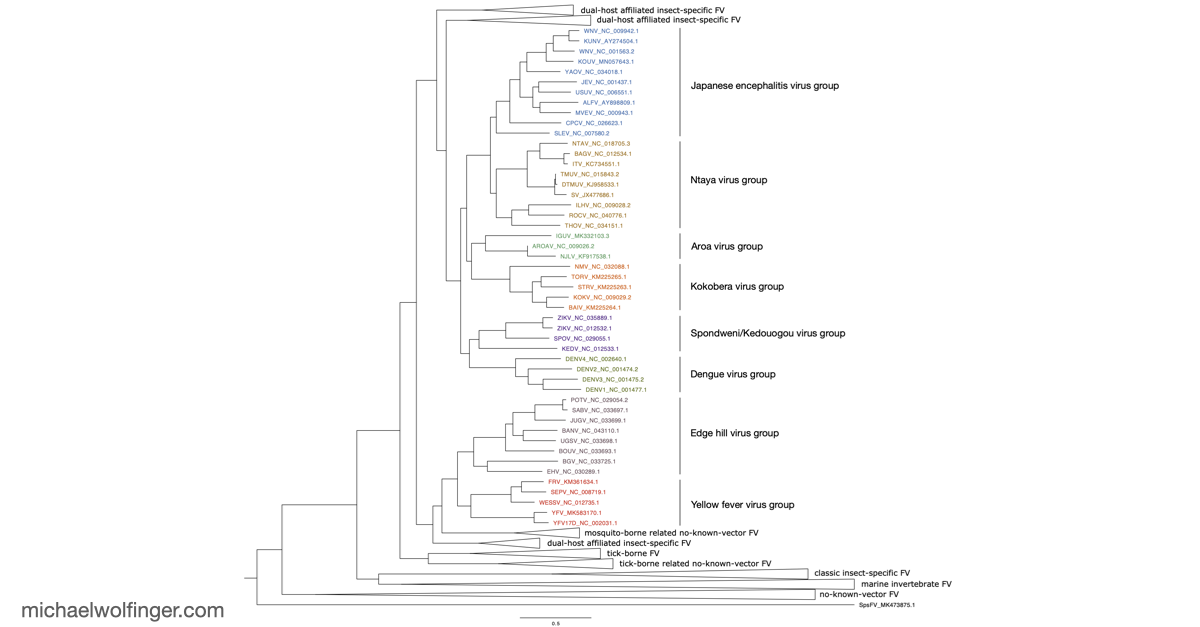
This chapter asks a deceptively simple question: what is actually conserved in the 3' UTRs of mosquito-borne flaviviruses once one looks beyond primary sequence? For viruses such as dengue, West Nile, yellow fever, Japanese encephalitis, and Zika virus, the answer matters because the structured 3' UTR is not passive genomic baggage. It is part of the replication program and helps shape genome cyclization, sfRNA production, host adaptation, and the broader regulatory behavior of the viral RNA.
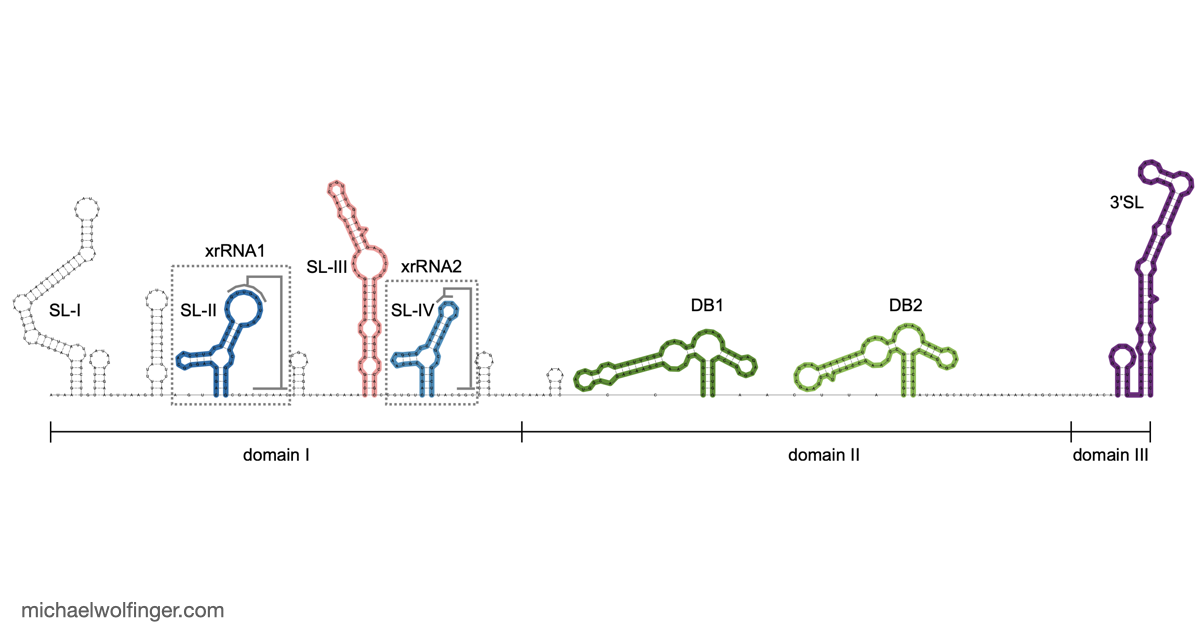
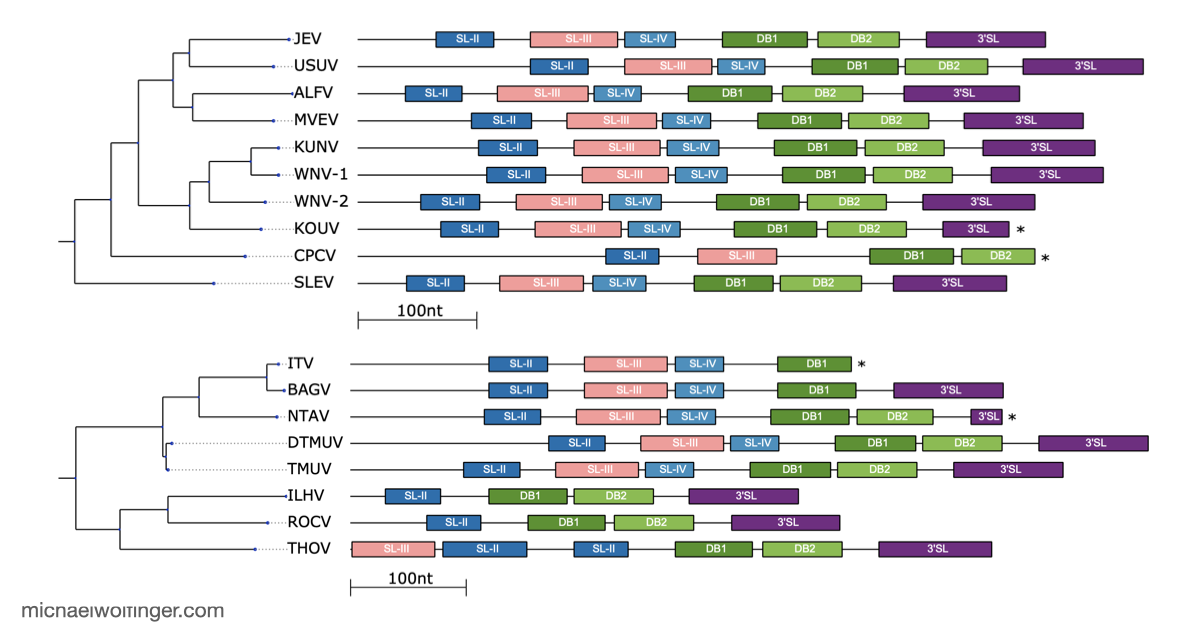
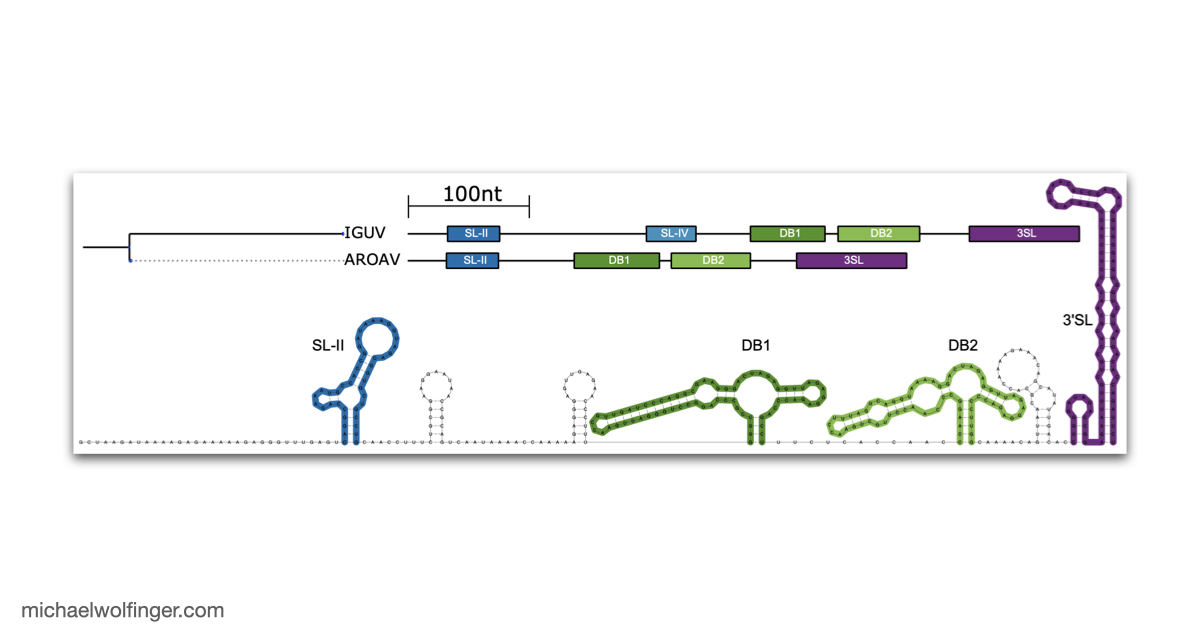
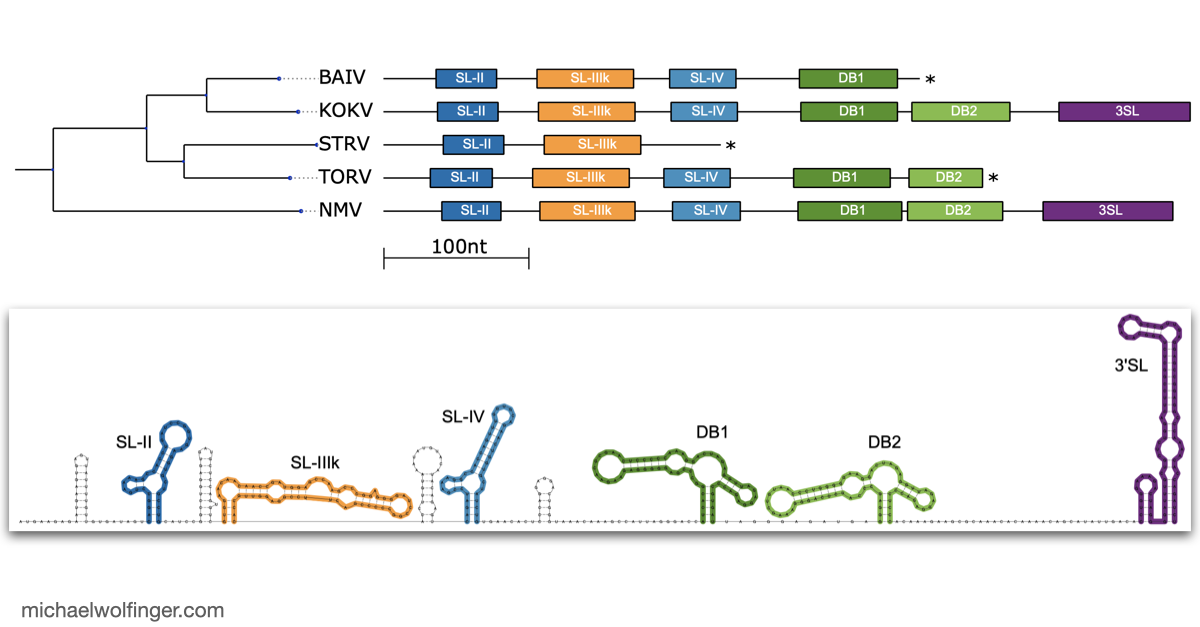
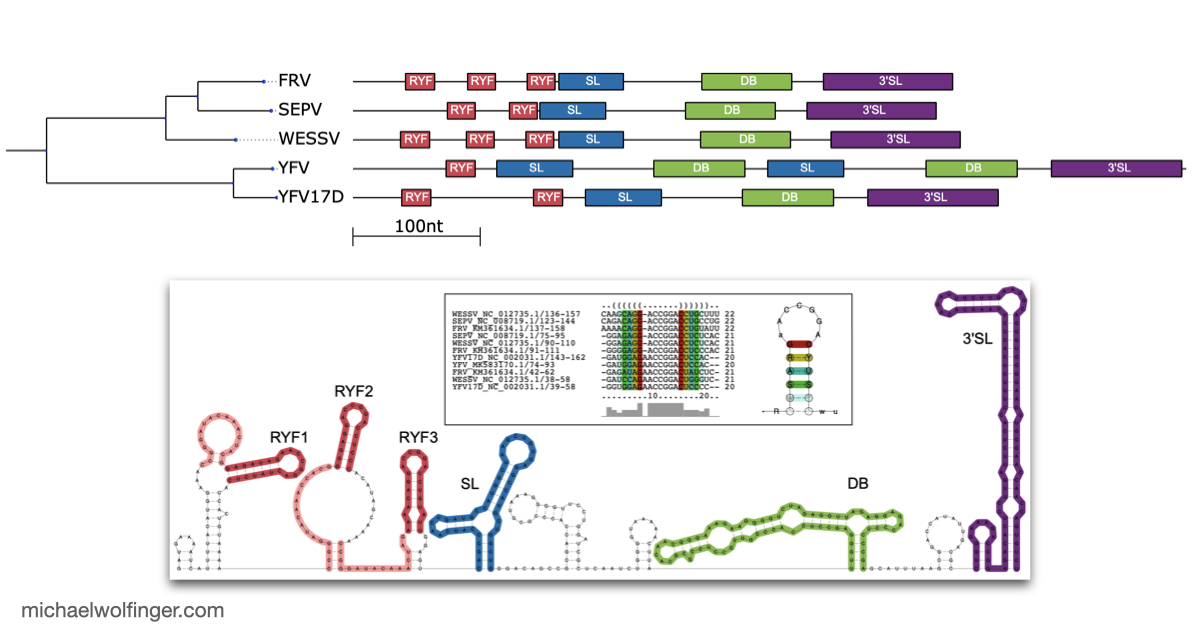
One useful starting point is that mosquito-borne flavivirus 3' UTRs do not consist of one canonical motif repeated from virus to virus. They are built from recurring structural modules, including xrRNAs, dumbbell elements, and terminal stem-loops, but the number, arrangement, and sequence realization of those modules differ across lineages. That means one has to think in terms of architectural conservation rather than exact sequence identity. The chapter pulls those patterns together in one place and shows where the common logic ends and lineage-specific variation begins.
The best known modules in this space are the exoribonuclease-resistant RNAs. xrRNAs stall host 5' to 3' nucleases such as Xrn1 and thereby protect downstream fragments that accumulate as subgenomic flaviviral RNAs. That mechanism has become one of the clearest examples of how RNA structure alone can create a biologically active molecular barrier. The chapter also makes the broader point that xrRNAs are only one part of the story. Mosquito-borne flaviviruses preserve other structured elements whose roles are tied to long-range RNA interactions, replication efficiency, and host-specific regulation.
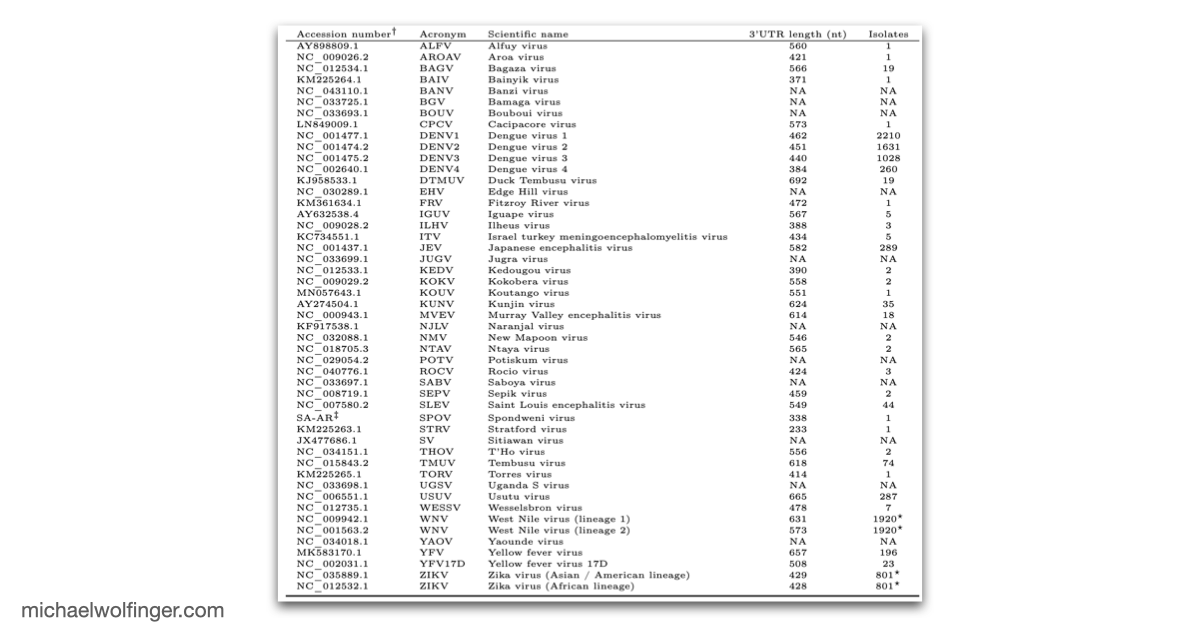
What makes the chapter valuable is the comparative scope. Instead of focusing on a single virus, it surveys the known mosquito-borne flavivirus diversity and asks which structural motifs recur across the group and which appear to have been reshaped during evolution. That perspective complements the earlier comparative 3' UTR analysis of tick-borne, insect-specific, and no-known-vector flaviviruses, where the emphasis was on broader flavivirus diversity and on the evolutionary distribution of xrRNA-like elements. Here the focus narrows to the mosquito-borne branch, where several of the best studied medically relevant flaviviruses sit.
That narrower focus also helps frame later, more specific papers. The logic of conserved 3' UTR architecture feeds directly into work on TBEV 3' UTR structure conservation and molecular epidemiology, as well as into experimental studies of structure-dependent functions such as the BVDV IRES analysis or later flavivirus papers on cyclization, G-quadruplexes, and small-RNA production. Once these untranslated regions are seen as structured regulatory platforms, many apparently separate virological questions start to line up more naturally.
For me, that is the real contribution of this chapter. It provides a map of the conserved structural vocabulary in mosquito-borne flavivirus 3' UTRs and makes it easier to ask which of those motifs are ancient, which are lineage-specific, and which are most likely to carry experimentally testable function. As a synthesis piece, it is less about a single new mechanistic claim than about organizing a rapidly expanding field into a coherent comparative picture.
That broader comparative point is also the theme of When sequence conservation is not enough to find functional RNA structure, which makes explicit why flaviviral 3'UTRs so often have to be read at the level of structural architecture rather than sequence similarity alone. Functional RNA Structures in the 3’UTR of Mosquito-Borne Flaviviruses Functional RNA Structures in the 3’UTR of Tick-Borne, Insect-Specific and No Known Vector Flaviviruses Evolutionary traits of Tick-borne encephalitis virus: Pervasive non-coding RNA structure conservation and molecular epidemiologyFigures and Data
Citation
Michael T. Wolfinger, Roman Ochsenreiter, Ivo L. Hofacker
In Virus Bioinformatics, edited by Dmitrij Frishman and Manja Marz, pp65–100. Chapman and Hall/CRC Press (2021) | doi: 10.1201/9781003097679-5 | PDF | FiguresSee Also
Roman Ochsenreiter, Ivo L. Hofacker, Michael T. Wolfinger
Viruses 11:298 (2019) | doi:10.3390/v11030298 | PDF | Figures
Lena S. Kutschera, Michael T. Wolfinger
Virus Evol. (8):1 veac051 (2022) | doi:10.1093/ve/veac051 | PDF | Figures