RNA structure conservation and molecular epidemiology of TBEV
This study combines comparative RNA structure analysis with molecular epidemiology to characterize conserved and variable 3' UTR architectures across tick-borne encephalitis virus lineages.
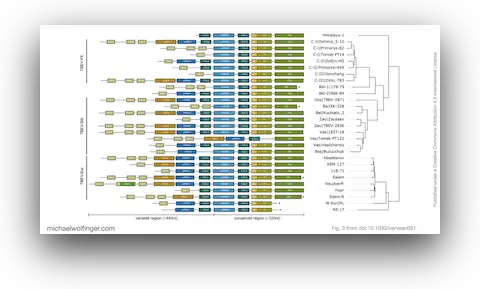
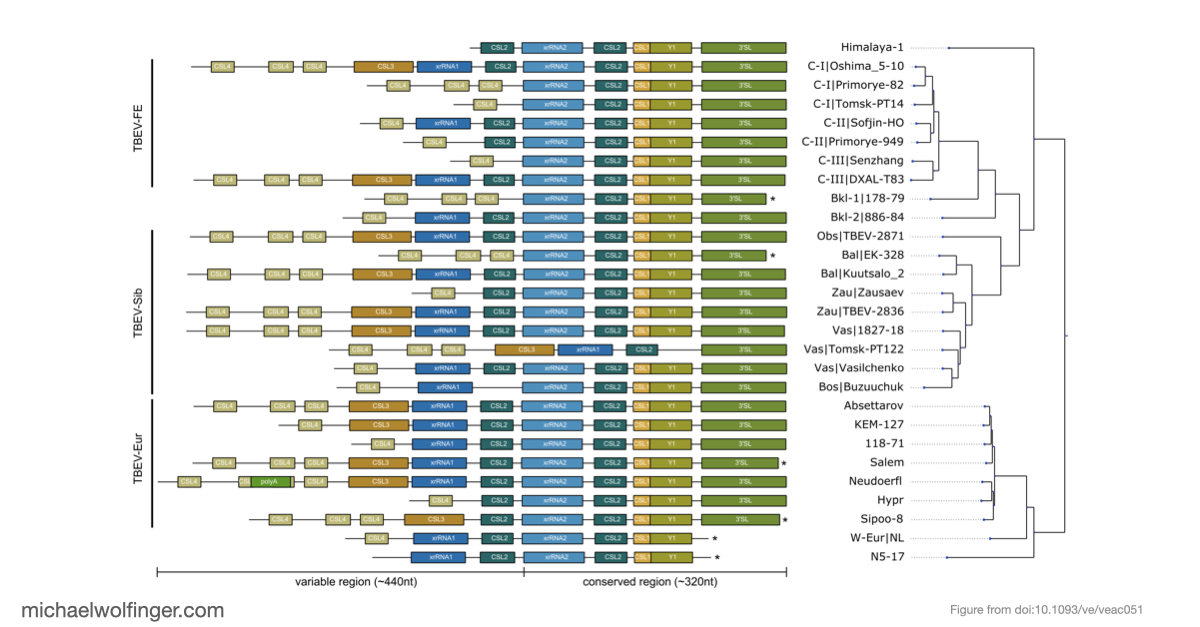
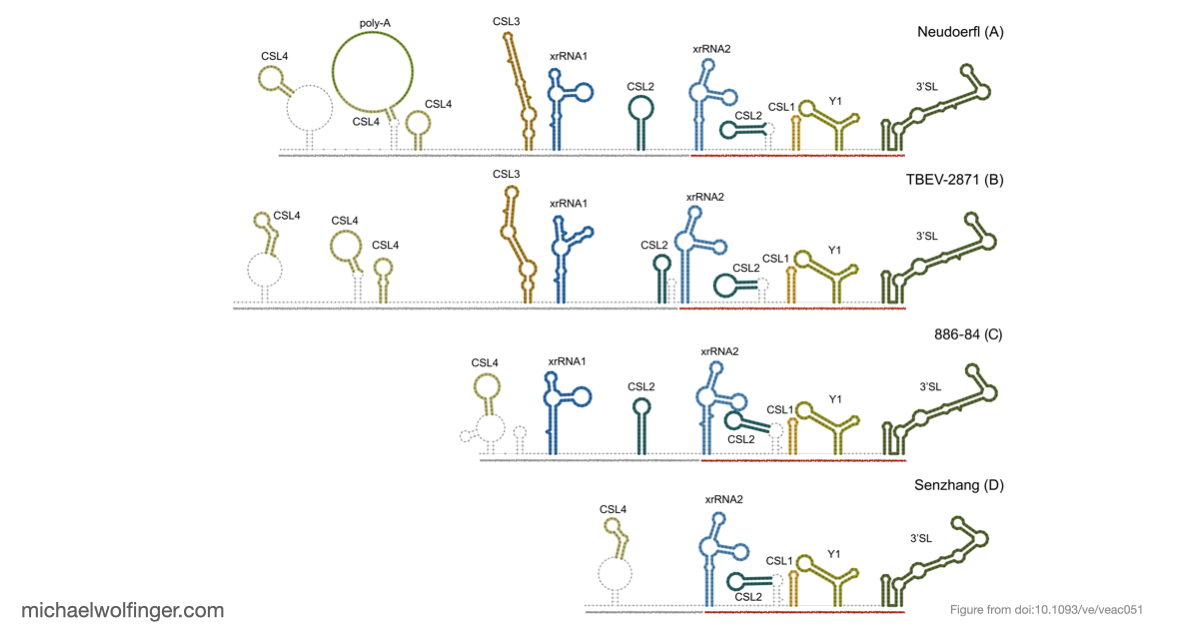
Tick-borne encephalitis virus is a good test case for comparative RNA virology because its 3' UTRs are variable enough to be evolutionarily interesting, but not so unconstrained that every lineage looks unrelated to the others. The paper asks whether that variation is best understood as random sequence drift or as remodeling within a limited structural vocabulary. The answer is clearly the latter. TBEV 3' UTRs contain a highly conserved core domain near the 3' end and a more labile upstream region, yet even the variable part can be described through recurring structured elements rather than unrestricted sequence turnover.
That point matters because it shifts the interpretation of TBEV diversity. If the architecture is built from a restricted set of conserved RNA modules, then lineage differences are not just a matter of "more" or "less" sequence divergence. They reflect different combinations, duplications, and losses of structured elements that are likely under functional constraint. In other words, the paper argues that 3' UTR evolution in TBEV is better understood at the level of RNA architecture than at the level of raw sequence identity alone.
The work fits naturally after the broader comparative flavivirus 3' UTR analysis and alongside the later mosquito-borne flavivirus 3' UTR synthesis. Those papers establish the larger structural vocabulary of flavivirus untranslated regions. This TBEV study then zooms in on one medically important lineage and asks how that vocabulary is reused within a single viral species complex. The narrower scope makes it easier to connect comparative RNA structure directly to questions of subtype diversification and geographic spread.
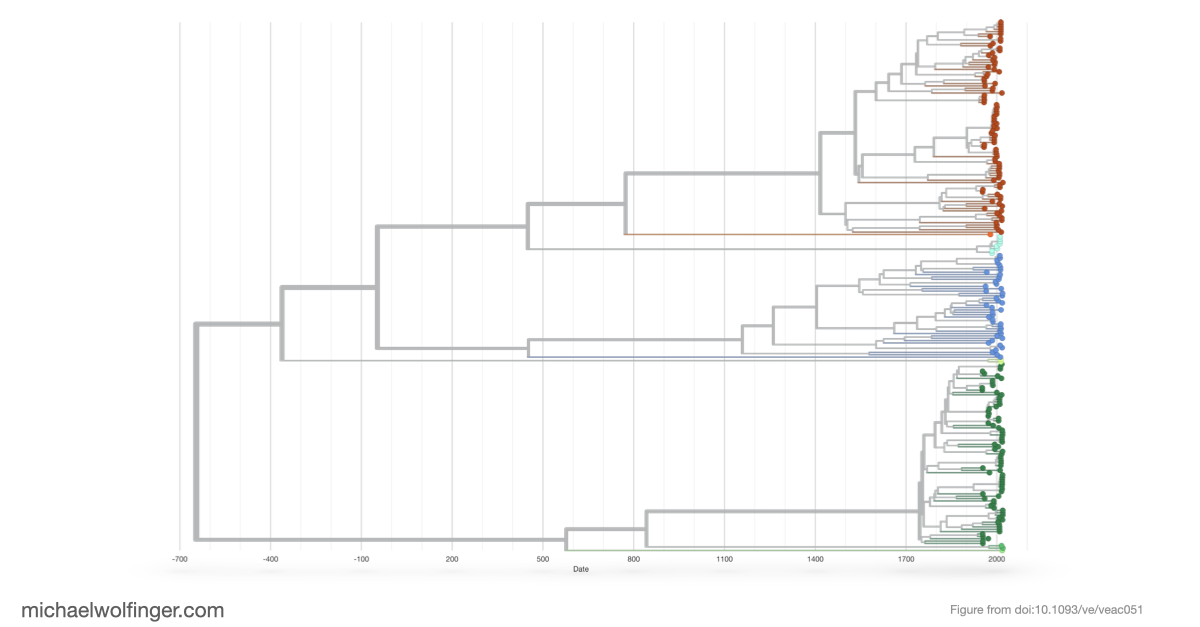
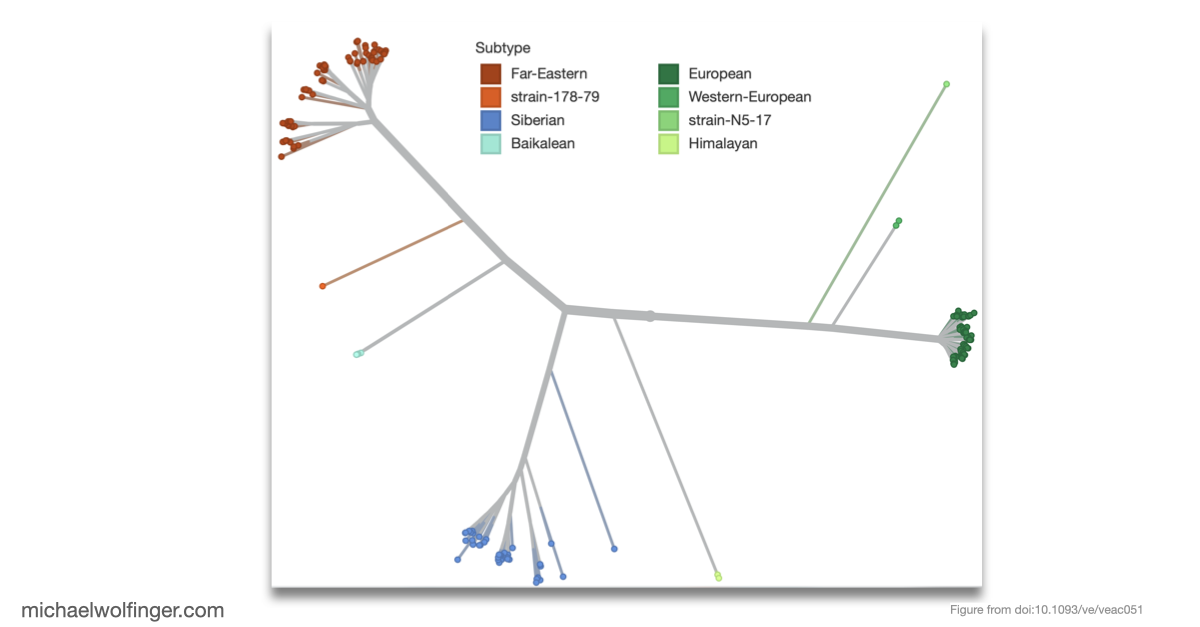
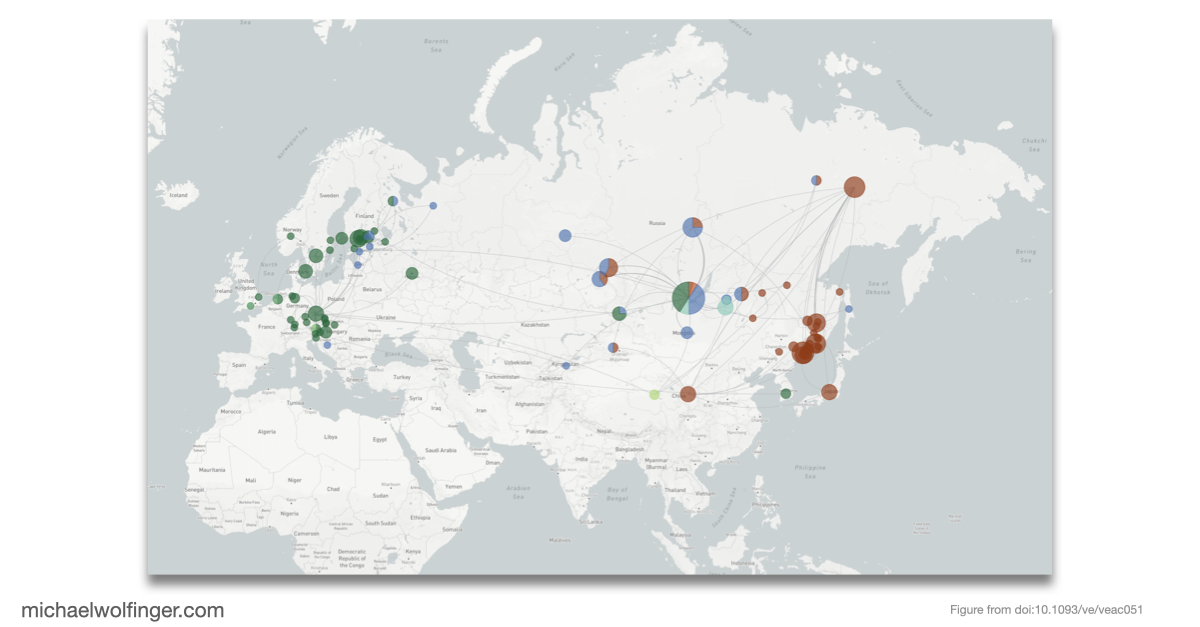
The molecular epidemiology part is therefore not an add-on. By introducing TBEVnext, the paper places the structural observations into an explicitly spatiotemporal framework. The resulting phylogeny shows that subtype labels and geographic occurrence do not map onto each other in a simple one-to-one way. That makes the dataset useful beyond visualization. It links particular 3' UTR architectures to the broader history of lineage expansion, ecological movement, and sampling across Europe and Asia.
What I find most compelling here is the combination of scales. The paper examines the fine-grained organization of non-coding RNA elements in the viral genome, then asks how those architectures sit inside the long-range history of the virus population. That is exactly what makes comparative RNA virology interesting. Structured RNA elements are not isolated curiosities. They evolve inside real lineages moving through hosts, vectors, and landscapes.
For TBEV specifically, the paper provides a more coherent picture of how structural conservation and epidemiological diversification coexist. The genome is not frozen, and the 3' UTR is certainly not uniform. Even so, the range of viable architectures appears constrained enough to reveal recurring principles. That makes the study a useful bridge between comparative structure prediction and phylodynamic surveillance, and gives the TBEV system a clearer place within the larger flavivirus RNA-structure landscape.
That broader point is exactly what I make more explicitly in When sequence conservation is not enough to find functional RNA structure. Evolutionary traits of Tick-borne encephalitis virus: Pervasive non-coding RNA structure conservation and molecular epidemiology Functional RNA Structures in the 3’UTR of Tick-Borne, Insect-Specific and No Known Vector Flaviviruses Functional RNA Structures in the 3’UTR of Mosquito-Borne FlavivirusesFigures and Data
Citation
Lena S. Kutschera, Michael T. Wolfinger
Virus Evol. (8):1 veac051 (2022) | doi: 10.1093/ve/veac051 | PDF | Supplementary dataSee Also
Roman Ochsenreiter, Ivo L. Hofacker, Michael T. Wolfinger
Viruses 11:298 (2019) | doi:10.3390/v11030298 | PDF | Figures
Michael T. Wolfinger, Roman Ochsenreiter, Ivo L. Hofacker
In Virus Bioinformatics, edited by Dmitrij Frishman and Manja Marz, pp65–100. Chapman and Hall/CRC Press (2021) | doi:10.1201/9781003097679-5 | Preprint PDF | Figures