Biophysical characterization of human lincRNA-p21 Alu inverted repeats
Biophysical and computational characterization of the tertiary structure of sense and antisense lincRNA-p21 Alu inverted repeats.
Human lincRNA-p21 is a regulatory long non-coding RNA in the p53 pathway and plays an important role in apoptosis. Its activity depends in part on its interaction with the RNA-binding protein hnRNP-K, and that interaction is likely shaped by RNA tertiary structure rather than sequence alone. Earlier work had already defined the secondary structure of the sense and antisense AluSx1 inverted repeats, but the three-dimensional organization of these RNA domains remained unclear.
This study closes that gap with an integrated biophysical and computational workflow. The sense and antisense AluSx1 regions were transcribed in vitro and first checked for sample quality and homogeneity using size-exclusion chromatography, analytical ultracentrifugation, and SEC-MALS. That part matters because long RNAs are structurally heterogeneous by default, and any useful SAXS model depends on showing that the sample is behaving as a monomeric full-length transcript rather than a mixture of fragments or aggregates.
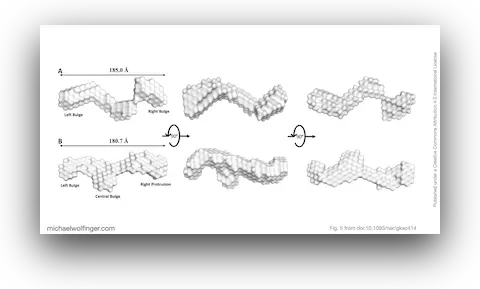
With that in place, the authors used SEC-SAXS to obtain low-resolution solution structures of both RNAs under near-physiological conditions. Both sense and antisense AluSx1 RNAs adopt elongated, asymmetric conformations rather than compact globular ones. The resulting SAXS envelopes suggest RNAs with extended double-stranded arms connected through junction regions, consistent with a structured but conformationally flexible architecture.
The computational part of the workflow then builds on previously determined secondary structures. Using SimRNA and hydrodynamic filtering, high-resolution atomistic models were generated and compared against the SAXS-derived envelopes. The important point is not that a single exact structure was identified, but that a constrained family of models can be found that agrees with both the biophysical data and the known secondary structure. In practice, this is a useful way to move from SHAPE-informed 2D models toward plausible 3D representations for long non-coding RNAs.
Biologically, that matters because the Alu inverted repeats are thought to contribute to how lincRNA-p21 interacts with protein partners and localizes within the cell. The study supports a picture in which both the sense and antisense regions form structured domains with conserved architectural features, while still retaining enough flexibility to make ensemble-based modeling more appropriate than a single rigid conformation.
More broadly, this paper is useful as a methods example. Long non-coding RNAs are difficult structural targets: they are large, flexible, and hard to crystallize, while sequence-only tertiary prediction remains unreliable. The pipeline used here, combining prior secondary-structure information with AUC, SEC-MALS, SAXS, and restrained computational modeling, is a practical template for deriving experimentally grounded structural models of other lncRNAs as well. Biophysical Characterisation of Human LincRNA-p21 Sense and Antisense Alu Inverted Repeats Michael H. D’Souza, Tyler Mrozowich, Maulik D. Badmalia, Mitchell Geeraert, Angela Frederickson, Amy Henrickson, Borries Demeler, Michael T. Wolfinger, Trushar R. PatelCitation
Nucleic Acids Res. gkac414 (2022) | doi:10.1093/nar/gkac414 | PDF