Hfq, Crc, and antibiotic resistance in P. aeruginosa
This study examines carbon catabolite repression and its indirect effects on antibiotic susceptibility in Pseudomonas aeruginosa.
This paper extends the Hfq/Crc/CrcZ story in Pseudomonas aeruginosa from single regulons to a dynamic physiological transition. The biological setting is diauxic growth: the bacterium first consumes a preferred carbon source and then rewires its metabolism once that source is depleted. In Pseudomonas, that transition is controlled in large part by carbon catabolite repression (CCR), with Hfq and Crc repressing many transcripts during growth on preferred substrates and the regulatory RNA CrcZ relieving that repression once the cell shifts to alternative nutrients.
The paper asks what that transition looks like globally and whether it has consequences beyond nutrient utilization. To answer that, the study measures the transcriptome, translatome, and proteome in parallel during and after relief of CCR. That multi-omics design is the real strength of the work. It allows the authors to distinguish changes that occur at RNA abundance, translational efficiency, and protein output, rather than treating "gene expression" as a single layer.
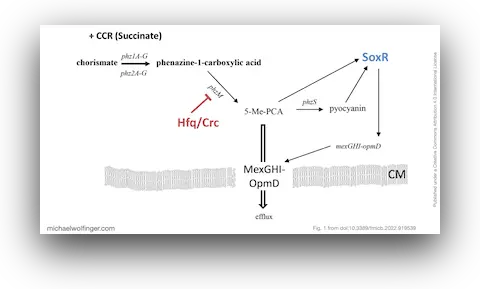
The main mechanistic result is that the mexGHI-opmD operon is upregulated after CCR is relieved, which in turn lowers susceptibility to norfloxacin. This is important because the operon encodes an efflux system with direct consequences for antibiotic response. The paper therefore shows that the shift from preferred to non-preferred carbon sources does not just alter metabolism. It also changes the antimicrobial phenotype of the cell.
That observation fits naturally into a broader line of work on how Hfq, Crc, and CrcZ link metabolism to RNA control in Pseudomonas. The mechanistic basis is laid out in Interplay Between the Catabolite Repression Control Protein Crc, Hfq and RNA in Hfq-Dependent Translational Regulation in Pseudomonas aeruginosa, and the link to carbapenem uptake is developed further in Distinctive Regulation of Carbapenem Susceptibility in Pseudomonas aeruginosa by Hfq. This paper pushes that logic one step further by showing how the same network indirectly reshapes resistance-relevant output during diauxic growth. Nutrient-state sensing and antibiotic susceptibility are tightly coupled through the same RNA-centered regulatory architecture.
The word "indirect" in the title matters. The study does not claim that Hfq binds the mexGHI-opmD operon in a simple one-step regulatory interaction. Instead, the data support a more distributed model in which relief of CCR changes the allocation and activity of Hfq-dependent control, and the effect on the efflux pump emerges from that broader rewiring. That makes the paper more interesting than a straightforward target-identification study, because it emphasizes network-level consequences of post-transcriptional regulation.
From a practical perspective, the result is also useful because it reminds us that antibiotic susceptibility can depend strongly on physiological state. The same bacterium can present a different drug-response profile depending on which nutrients it has consumed and which regulatory program it has entered. That is exactly the kind of context dependence that often complicates antimicrobial treatment and laboratory interpretation.
Methodologically, the paper is a good example of how multi-omics becomes genuinely informative when tied to a clear transition state. Sampling during and after relief of CCR gives the authors a biologically meaningful perturbation, and the combined transcriptome-translatome-proteome view makes it possible to see which responses are broad and which are more specifically post-transcriptional. For readers interested in bacterial RNA regulation, that is a major part of the value.
Taken together with the 2018 and 2020 studies, this paper makes the connection between metabolism, RNA control, and resistance phenotypes especially clear. It shows how the same regulatory machinery is deployed during a physiological growth transition and how that deployment feeds into efflux-mediated drug response. Rewiring of Gene Expression in Pseudomonas aeruginosa During Diauxic Growth Reveals an Indirect Regulation of the MexGHI-OpmD Efflux Pump by HfqCitation
Marlena Rozner, Ella Nukarinen, Michael T. Wolfinger, Fabian Amman, Wolfram Weckwerth, Udo Blaesi, Elisabeth Sonnleitner
Front. Microbiol. (2022) 13:919539 | doi:10.3389/fmicb.2022.919539 | PDF