Mpulungu virus and unique xrRNAs in a novel African tick flavivirus
This study describes Mpulungu virus, a novel African tick flavivirus, and characterizes unusual exoribonuclease-resistant RNA structures in its 3' UTR.
Mpulungu virus became important because it did not fit comfortably into the usual picture of tick-borne flaviviruses. The genome recovered from a Rhipicephalus tick in Zambia grouped with Ngoye virus from Senegal, defining an unusual African lineage that sat apart from the better known vertebrate-associated tick-borne flaviviruses. At the time, that alone made the paper notable. It expanded the geographic and phylogenetic range of the group and suggested that the ecological diversity of tick-associated flaviviruses had been underestimated.
What gives the paper lasting value is the structural analysis of the untranslated regions. The 5' end of MPFV still looks recognizably flaviviral, but the 3' UTR is where the more interesting divergence appears. Instead of treating the non-coding region as an unusual sequence tail, the paper asks which structured RNA elements are preserved there and whether they remain functionally competent despite the lineage's broader divergence.
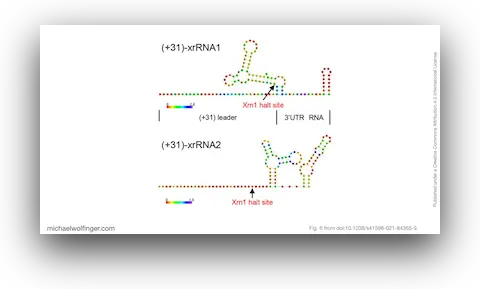
The central result is the identification of two exoribonuclease-resistant RNA elements in the 3' UTR of MPFV. These xrRNA structures are not just predicted computationally. The study shows biochemically that both can stall Xrn1, providing direct evidence that this unusual African lineage retains a key flaviviral strategy for generating protected decay intermediates and, likely, subgenomic flaviviral RNAs. That tied the new lineage back to a conserved mechanistic theme in flavivirus RNA biology even while the surrounding genomic context looked quite distinctive.
Seen this way, MPFV was an early sign that unusual tick-associated flaviviruses could preserve the broader flaviviral RNA toolkit while following different ecological trajectories. The later Xinyang flavivirus paper sharpened that interpretation by showing that the lineage was not confined to Africa and by strengthening the case for a basal, likely tick-only clade. Read together, the two papers make more sense than either one does alone. MPFV established the branch and its unusual xrRNA features. XiFV then broadened the clade and shifted the ecological argument away from a vertebrate-centered default.
This also places MPFV naturally beside the Zambian insect-specific flavivirus xrRNA study. In both cases, virus discovery becomes much more informative once it is connected to comparative RNA structure and functional validation. The broader lesson is that understudied flavivirus lineages often reveal their deepest commonalities not through obvious sequence identity, but through preserved structured RNA elements in the untranslated regions.
I would now read this paper less as a warning about an immediate public-health threat and more as a foundational comparative RNA virology study. It identified a novel lineage, mapped its structured 3' UTR, and showed that even in a phylogenetically unusual branch, xrRNA function remains central enough to be retained. That is the kind of result that makes later ecological and evolutionary reinterpretation possible.
It is also a good example of the argument in When sequence conservation is not enough to find functional RNA structure, because the deeper commonality here emerges from preserved structured-RNA logic rather than from easy sequence similarity alone. An African Tick Flavivirus Forming an Independent Clade Exhibits Unique Exoribonuclease-Resistant RNA Structures in the Genomic 3’-Untranslated Region Xinyang flavivirus, from Haemaphysalis flava ticks in Henan province, China, defines a basal, likely tick-only flavivirus cladeCitation
Hayato Harima, Yasuko Orba, Shiho Torii, Yongjin Qiu, Masahiro Kajihara, Yoshiki Eto, Naoya Matsuta, Bernard M. Hang’ombe, Yuki Eshita, Kentaro Uemura, Keita Matsuno, Michihito Sasaki, Kentaro Yoshii, Ryo Nakao, William W. Hall, Ayato Takada, Takashi Abe, Michael T. Wolfinger, Martin Simuunza, Hirofumi Sawa
Sci. Rep. 11:4883 (2021) | doi: 10.1038/s41598-021-84365-9 | PDFSee Also
Lan-Lan Wang, Qia Cheng, Natalee D. Newton, Michael T. Wolfinger, Mahali S. Morgan, Andrii Slonchak, Alexander A. Khromykh, Tian-Yin Cheng, Rhys H. Parry
J. Gen. Virol. 105(5) (2024) | doi:10.1099/jgv.0.001991 | PDF