Musashi binding elements in Zika virus 3'UTR
A comparative analysis of Musashi binding element accessibility in Zika virus and related flavivirus 3' UTRs using thermodynamic RNA structure modeling.
Musashi-1 is an RNA-binding protein that is highly expressed in neural stem and progenitor cells, and earlier experimental work had suggested that it can bind the Zika virus 3' UTR and enhance viral replication. That makes the structural context of Musashi binding elements in viral RNA immediately relevant: binding depends not just on the presence of a UAG core motif, but on whether that motif is exposed in a single-stranded region that is accessible to the protein.
This paper addresses exactly that question with a thermodynamic RNA-structure model. We screened curated flavivirus 3' UTRs for UAG motifs and evaluated how accessible each site is in its native sequence context. Rather than just folding a single minimum free energy structure, the analysis uses opening energies derived from ensemble calculations in sliding windows and compares them to dinucleotide-shuffled sequence controls. The resulting z scores make it possible to ask whether a given Musashi binding element is unusually exposed relative to what would be expected from local sequence composition alone.
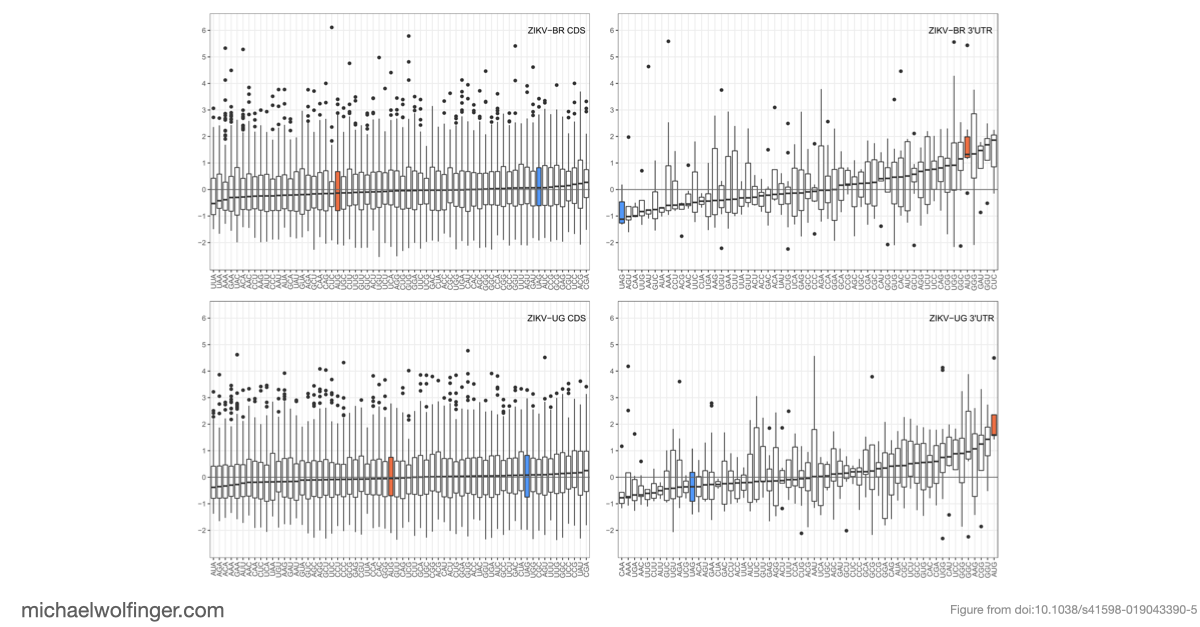
For Zika virus, the result is clear. In both the African and the Asian/American lineages, UAG belongs to the most accessible trinucleotides in the 3' UTR, and the surrounding pentanucleotide context remains accessible as well. The Asian/American lineage is especially striking: in the Brazilian isolate used here, all UAG motifs in the 3' UTR show negative opening-energy z scores, meaning that they are consistently predicted to occur in unpaired structural contexts. That fits well with the idea that Musashi binding is structurally favored in epidemic ZIKV strains.
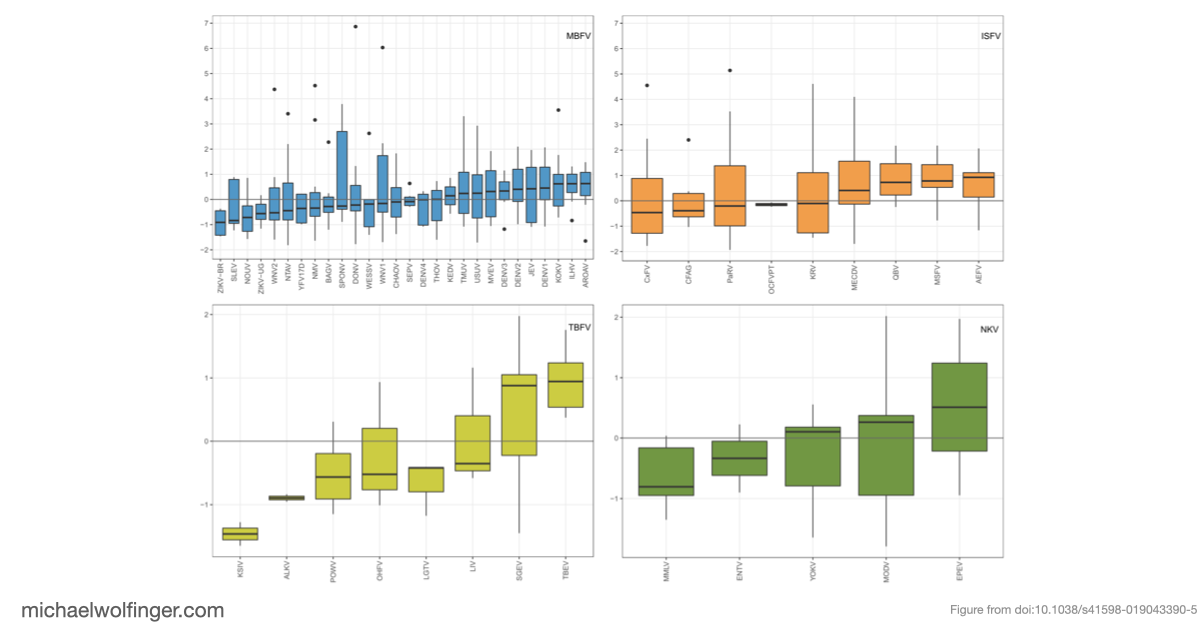
The comparative part of the study is just as important. The same calculation was carried out across mosquito-borne, tick-borne, insect-specific, and no-known-vector flaviviruses. ZIKV from the Asian/American lineage ranked as the most Musashi-accessible among the mosquito-borne flaviviruses in the dataset, but it was not the only virus with accessible Musashi motifs. Some West Nile, yellow fever, Powassan, Karshi, and insect-specific flaviviruses also contain strongly accessible UAG sites. That does not mean that all of these viruses share the same neuropathology as ZIKV, but it does show that ZIKV is not uniquely equipped with Musashi-compatible RNA motifs.
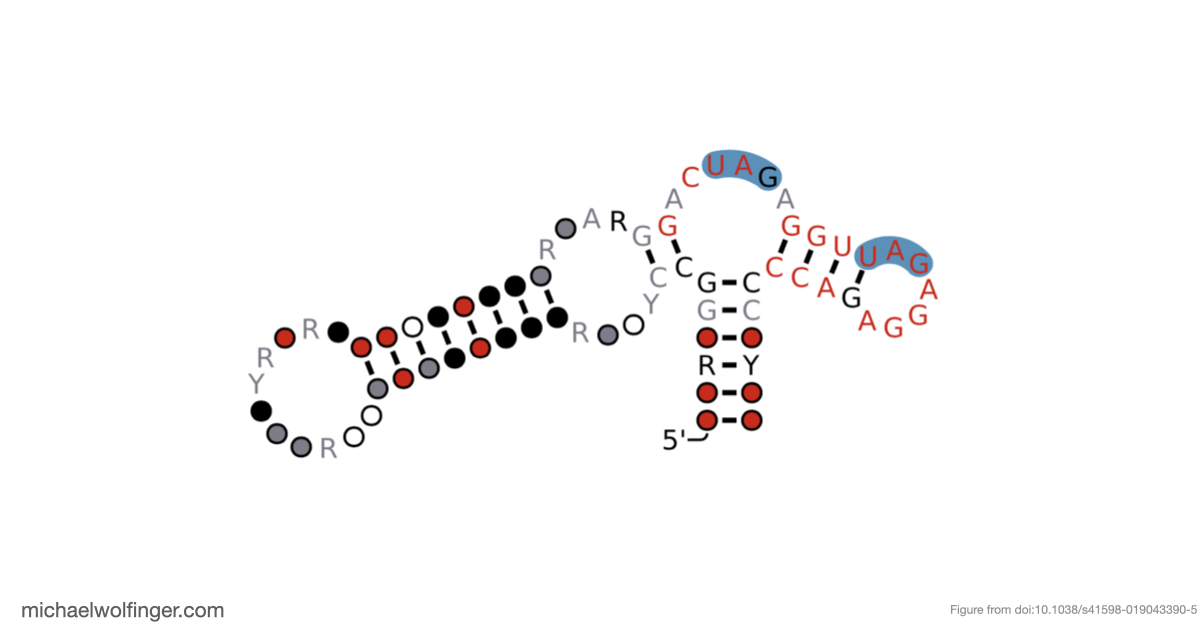
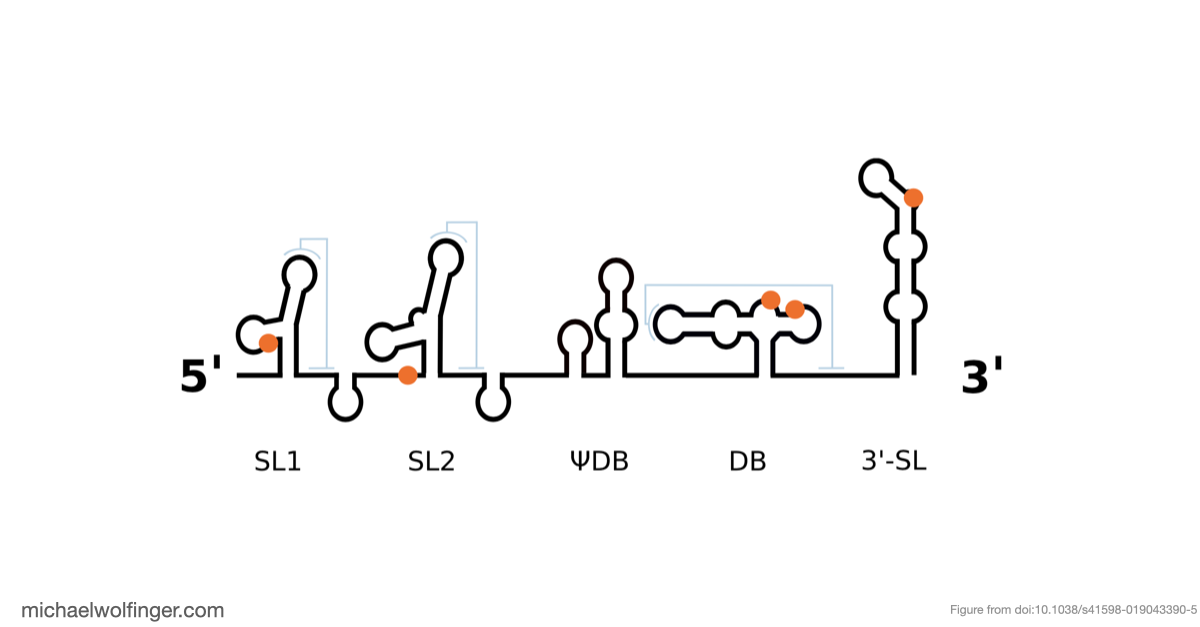
Another useful result is that Musashi binding elements are not randomly scattered across flavivirus 3' UTRs. By mapping the motifs onto conserved structural elements, the study found that dumbbell elements in particular often carry two conserved UAG motifs in a shared structural context. That is interesting because dumbbell RNAs are already known as functionally important 3' UTR elements in many flaviviruses. The combination of structural conservation and repeated Musashi-compatible motifs suggests that these sites are worth testing experimentally rather than treating them as incidental sequence matches.
Conceptually, this work is less about proving a single mechanism and more about building a tractable computational screen for host-factor compatibility in viral RNAs. Accessibility alone is not enough to predict congenital infection or neurotropism, and the paper is careful not to claim that it is. But opening-energy calculations do provide a useful way to prioritize candidate binding sites and candidate viruses for follow-up experiments. In that sense, the study connects RNA secondary structure thermodynamics with a concrete virological question: which flaviviruses place Musashi binding motifs in structural contexts that are likely to matter biologically? Musashi Binding Elements in Zika and Related Flavivirus 3’UTRs: A Comparative Study in Silico Theoretical studies on RNA recognition by Musashi 1 RNA–binding protein A Structural Refinement Technique for Protein-RNA Complexes Using a Combination of AI-based Modeling and Flexible Docking: A Study of Musashi-1 ProteinFigures and Data
Citation
Adriano de Bernardi Schneider, Michael T. Wolfinger
Sci. Rep. 9(1):6911 (2019) | doi:10.1038/s41598-019-43390-5 | PDF | FiguresSee Also
Nitchakan Darai, Panupong Mahalapbutr, Peter Wolschann, Vannajan Sanghiran Lee, Michael T. Wolfinger, Thanyada Rungrotmongkol
Sci. Rep. 12:12137 (2022) | doi:10.1038/s41598-022-16252-w | PDF | Figures
Nitchakan Darai, Kowit Hengphasatporn, Peter Wolschann, Michael T. Wolfinger, Yasuteru Shigeta, Thanyada Rungrotmongkol, Ryuhei Harada
B. Chem. Soc. Jpn. 96(7):677–685 (2023) | doi:10.1246/bcsj.20230092 | PDF