RNA recognition by Musashi-1
Molecular dynamics and binding-energy calculations are used here to compare how Musashi-1 recognizes different RNA motifs and to identify determinants of binding specificity.
Musashi-1 (MSI1) is an RNA-binding protein involved in stem-cell maintenance, neural development, and post-transcriptional regulation. It also became particularly interesting in the context of Zika virus biology, because Musashi proteins were proposed to interact with viral RNAs in ways that could affect replication and neuropathology. That makes the underlying recognition problem more than a narrow structural question. Understanding how Musashi binds RNA is relevant both for endogenous regulation and for virus-host interaction studies.
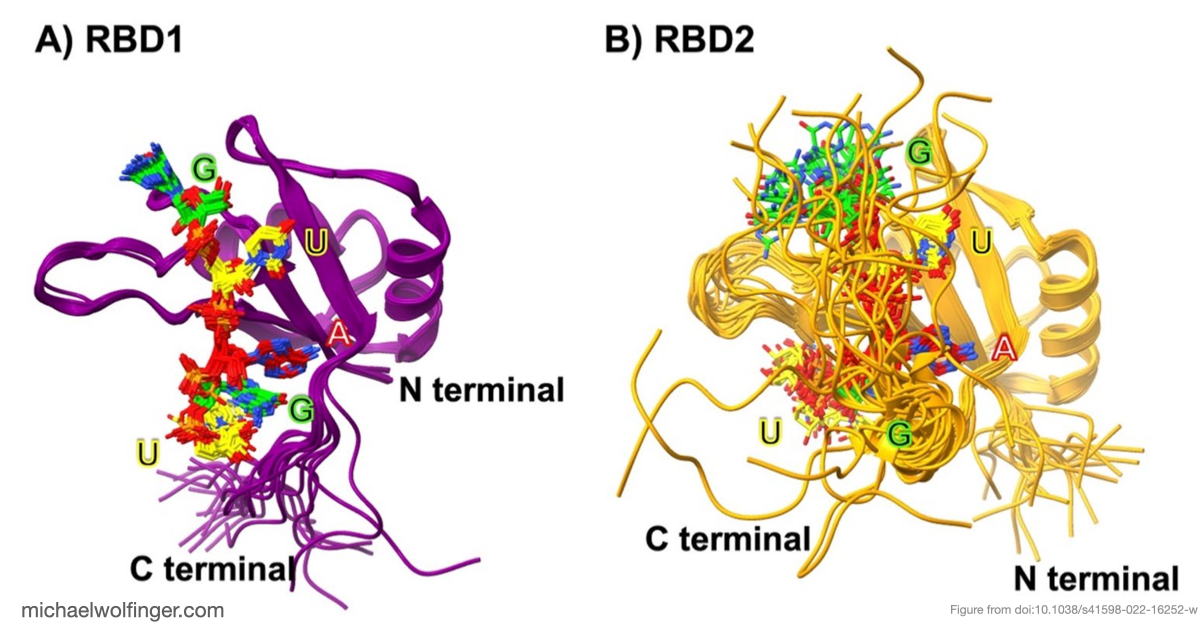
This paper focuses on that recognition step at the level of individual binding domains. MSI1 contains two RNA-binding domains, RBD1 and RBD2, both of which recognize short single-stranded RNA motifs centered on a UAG core. The main question is how specific that recognition really is. Sequence logos and motif descriptions are useful, but they do not explain why some candidate pentamers bind more strongly than others or which contacts stabilize the preferred complexes.
The methodological approach is straightforward but well chosen for this kind of problem. The study starts from structural models for the Musashi RNA-binding domains and combines those with several candidate RNA pentamers. These protein-RNA complexes are then evaluated using atomistic molecular dynamics simulations in explicit solvent, followed by binding free-energy estimates using the solvated interaction energy framework. In practice, that means the paper does not rely on a single static docking pose. It asks whether the complexes remain stable over time and whether the interaction energies consistently favor some RNA motifs over others.
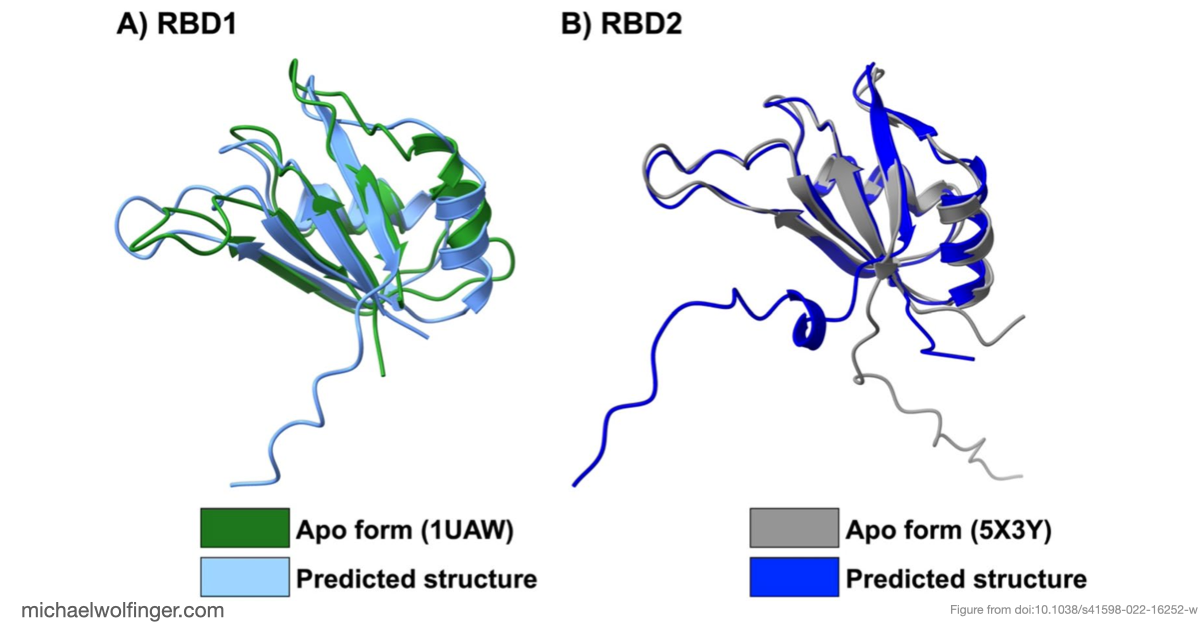
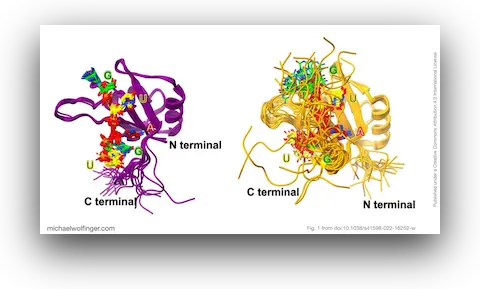
An additional point of interest is the use of AlphaFold2 to evaluate the protein side of the system. At the time, this was a useful check on whether predicted Musashi domain structures were close enough to experimentally determined conformations to support downstream simulation. The answer was essentially yes. The predicted structures aligned well with known domain architectures, which made it reasonable to use them as a basis for analyzing RNA recognition.
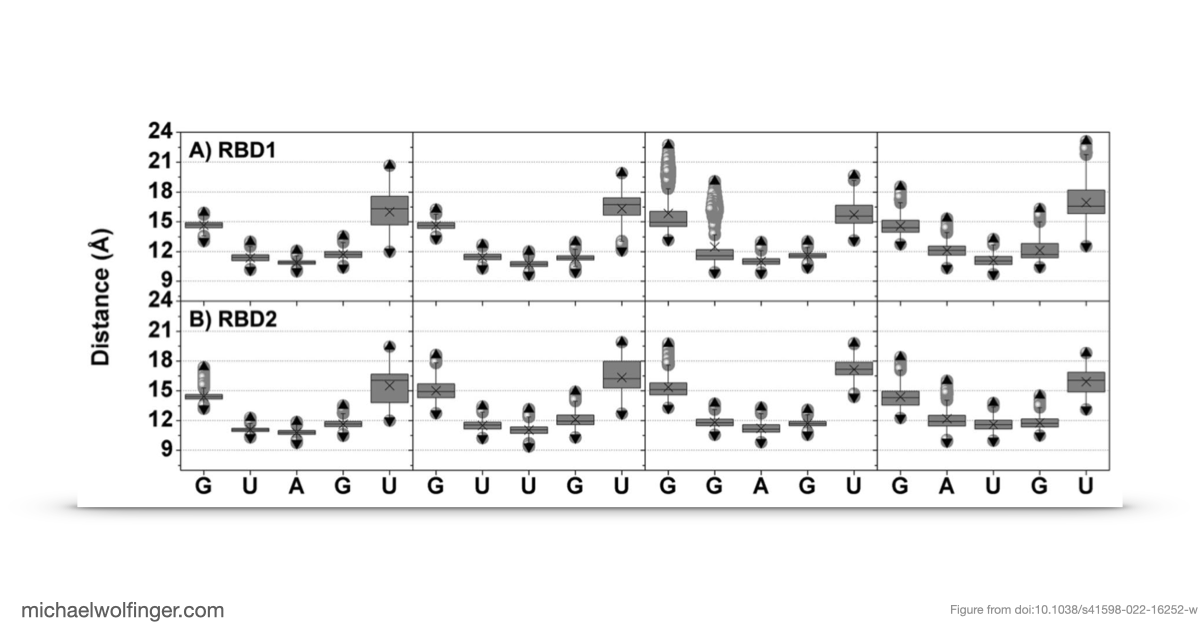
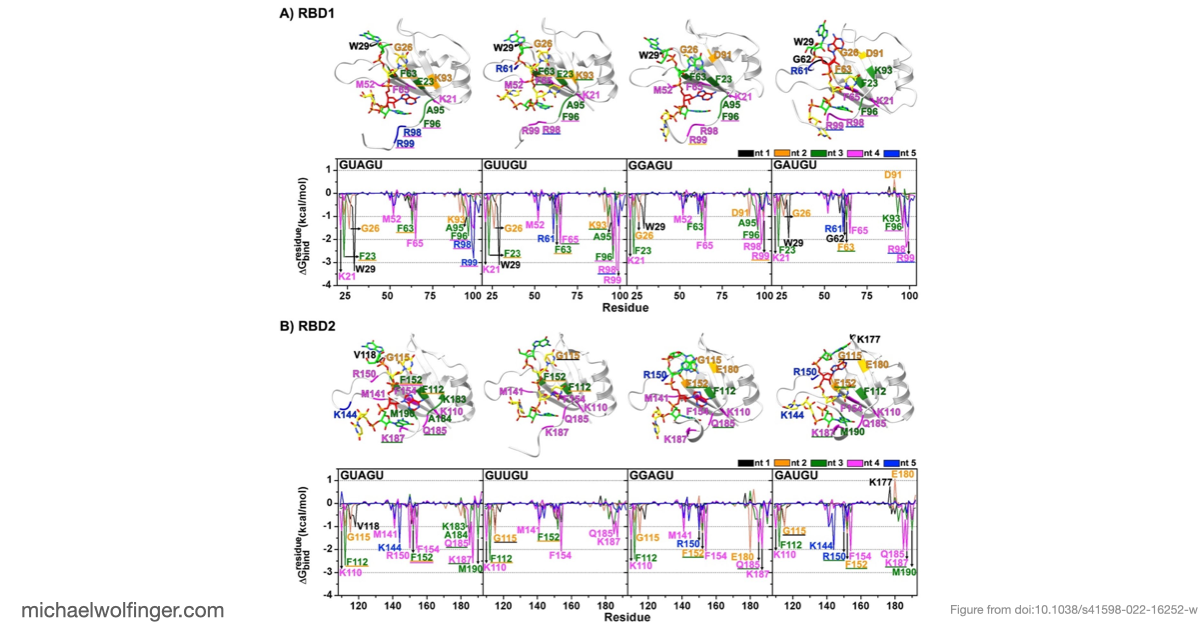
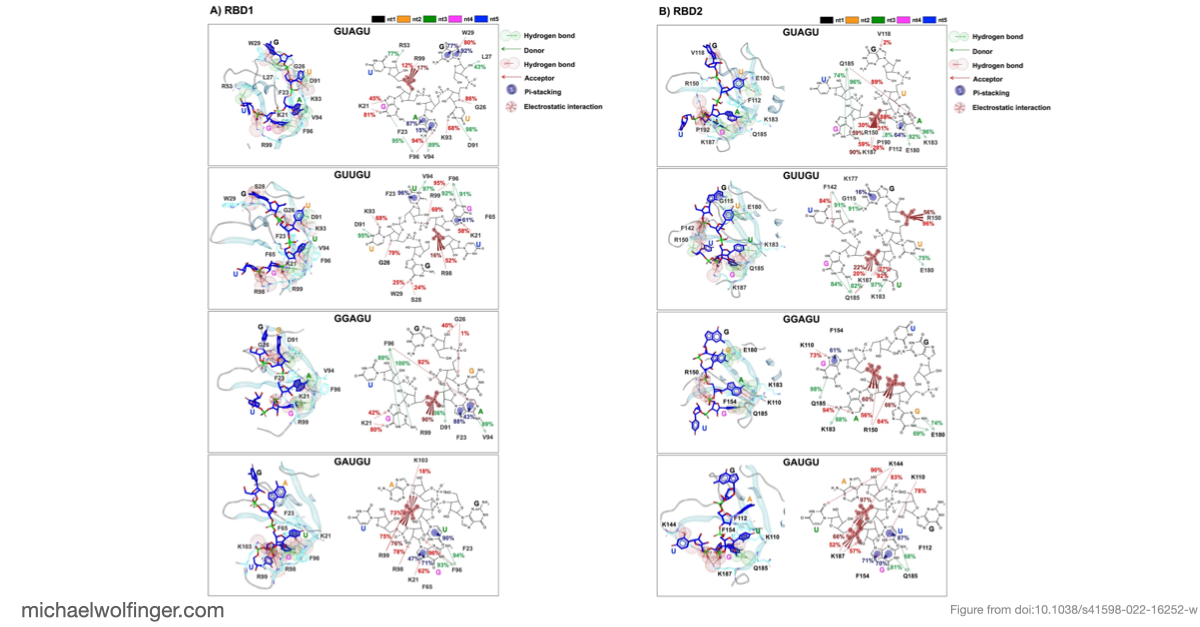
The main finding is that Musashi binding is selective even among short, closely related RNA pentamers. Out of the four tested motifs, two showed substantially stronger calculated binding to MSI1 RBD1 and RBD2 than the others. The simulations trace that difference back to a more favorable network of stacking, hydrogen-bonding, and electrostatic interactions around the central recognition motif. The work moves the discussion from "Musashi prefers UAG-containing RNAs" to a more specific structural claim about why some local sequence contexts are better accommodated than others.
That is what makes the paper useful. It does not claim to solve RNA-protein recognition in general, and it does not replace experiment. It does provide a mechanistic picture of how Musashi domains engage short RNA targets and how binding specificity can be compared systematically across candidate motifs. For systems where direct structural data are still incomplete, that kind of analysis is often the right intermediate step between motif discovery and functional testing.
This study also laid some groundwork for later Musashi modeling efforts. Once domain-level recognition preferences are characterized at this level, it becomes easier to ask larger questions about Musashi binding to structured cellular RNAs or to viral untranslated regions, and to develop more detailed protein-RNA complex refinement workflows. In that sense, this paper is best viewed as the starting point of a broader Musashi-RNA modeling program rather than a standalone binding-energy exercise.
For a broader view of how AI enters protein-RNA modeling beyond this individual Musashi case, see What AI can and cannot do for RNA structure and RNA-protein modeling. Theoretical studies on RNA recognition by Musashi 1 RNA–binding protein Musashi Binding Elements in Zika and Related Flavivirus 3’UTRs: A Comparative Study in Silico A Structural Refinement Technique for Protein-RNA Complexes Using a Combination of AI-based Modeling and Flexible Docking: A Study of Musashi-1 ProteinFigures and Data
Citation
Nitchakan Darai, Panupong Mahalapbutr, Peter Wolschann, Vannajan Sanghiran Lee, Michael T. Wolfinger, Thanyada Rungrotmongkol
Sci. Rep. 12:12137 (2022) | doi:10.1038/s41598-022-16252-w | PDFSee Also
Adriano de Bernardi Schneider, Michael T. Wolfinger
Sci. Rep. 9(1):6911 (2019) | doi:10.1038/s41598-019-43390-5 | PDF | Figures
Nitchakan Darai, Kowit Hengphasatporn, Peter Wolschann, Michael T. Wolfinger, Yasuteru Shigeta, Thanyada Rungrotmongkol, Ryuhei Harada
B. Chem. Soc. Jpn. 96(7):677–685 (2023) | doi:10.1246/bcsj.20230092 | PDF